Home » Posts tagged 'genes'
Tag Archives: genes
DNA—Blueprint and Fortune Teller?
2500 words
What would you think if you heard about a new fortune-telling device that is touted to predict psychological traits like depression, schizophrenia and school achievement? What’s more, it can tell your fortune from the moment of your birth, it is completely reliable and unbiased — and it only costs £100.
This might sound like yet another pop-psychology claim about gimmicks that will change your life, but this one is in fact based on the best science of our times. The fortune teller is DNA. The ability of DNA to understand who we are, and predict who we will become has emerged in the last three years, thanks to the rise of personal genomics. We will see how the DNA revolution has made DNA personal by giving us the power to predict our psychological strengths and weaknesses from birth. This is a game-changer as it has far-reaching implications for psychology, for society and for each and every one of us.
This DNA fortune teller is the culmination of a century of genetic research investigating what makes us who we are. When psychology emerged as a science in the early twentieth century, it focused on environmental causes of behavior. Environmentalism — the view that we are what we learn — dominated psychology for decades. From Freud onwards, the family environment, or nurture, was assumed to be the key factor in determining who we are. (Plomin, 2018: 6, my emphasis)
The main premise of Plomin’s 2018 book Blueprint is that DNA is a fortune teller while personal genomics is a fortune-telling device. The fortune-telling device Plomin most discusses in the book is polygenic scores (PGS). PGSs are gleaned from GWA studies; SNP genotypes are then added up with scores of 0, 1, and 2. Then, the individual gets their PGS for trait T. Plomin’s claim—that DNA is a fortune teller—though, falls since DNA is not a blueprint—which is where the claim that “DNA is a fortune teller” is derived.
It’s funny that Plomin calls the measure “unbiased”, (he is talking about DNA, which is in effect “unbiased”), but PGS are anything BUT unbiased. For example, most GWAS/PGS are derived from European populations. But, for example, there are “biases and inaccuracies of polygenic risk scores (PRS) when predicting disease risk in individuals from populations other than those used in their derivation” (De La Vega and Bustamante, 2018). (PRSs are derived from statistical gene associations using GWAS; Janssens and Joyner, 2019.) Europeans make up more than 80 percent of GWAS studies. This is why, due to the large amount of GWASs on European populations, that “prediction accuracy [is] reduced by approximately 2- to 5-fold in East Asian and African American populations, respectively” (Martin et al, 2018). See for example Figure 1 from Martin et al (2018):
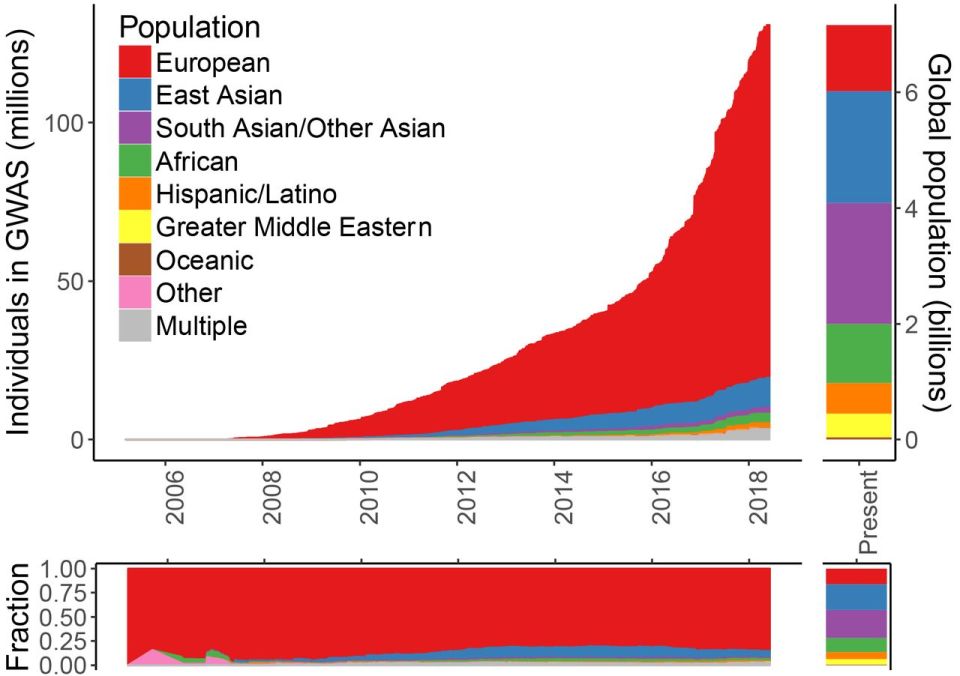
With the huge number of GWAS studies done on European populations, these scores cannot be used on non-European populations for ‘prediction’—even disregarding the other problems with PGS/GWAS.
By studying genetically informative cases like twins and adoptees, behavioural geneticists discovered some of the biggest findings in psychology because, for the first time, nature and nurture could be disentangled.
[…]
… DNA differences inherited from our parents at the moment of conception are the consistent, lifelong source of psychological individuality, the blueprint that makes us who we are. A blueprint is a plan. … A blueprint isn’t all that matters but it matters more than everything else put together in terms of the stable psychological traits that make us who we are. (Plomin, 2018: 6-8, my emphasis)
Nevermind the slew of problems with twin and adoption studies (Joseph, 2014; Joseph et al, 2015; Richardson, 2017a). I also refuted the notion that “A blueprint is a plan” last year, quoting numerous developmental systems theorists. The main thrust of Plomin’s book—that DNA is a blueprint and therefore can be seen as a fortune teller using the fortune-telling device to tell the fortunes of the people’s whose DNA are analyzed—is false, as DNA does not work how it does in Plomin’s mind.
These big findings were based on twin and adoption studies that indirectly assessed genetic impact. Twenty years ago the DNA revolution began with the sequencing of the human genome, which identified each of the 3 billion steps in the double helix of DNA. We are the same as every other human being for more than 99 percent of these DNA steps, which is the blueprint for human nature. The less than 1 per cent of difference of these DNA steps that differ between us is what makes us who we are as individuals — our mental illnesses, our personalities and our mental abilities. These inherited DNA differences are the blueprint for our individuality …
[DNA predictors] are unique in psychology because they do not change during our lives. This means that they can foretell our futures from our birth.
[…]
The applications and implications of DNA predictors will be controversial. Although we will examine some of these concerns, I am unabashedly a cheerleader for these changes. (Plomin, 2018: 8-10, my emphasis)
This quote further shows Plomin’s “blueprint” for the rest of his book—DNA can “foretell our futures from our birth”—and how it affects his conclusions gleaned from his work that he mostly discusses in his book. Yes, all scientists are biased (as Stephen Jay Gould noted), but Plomin outright claimed to be an unabashed cheerleader for his work. Plomin’s self-admission for being an “unabashed cheerleader”, though, does explain some of the conclusions he makes in Blueprint.
However, the problem with the mantra ‘nature and nurture’ is that it runs the risk of sliding back into the mistaken view that the effects of genes and environment cannot be disentangled.
[…]
Our future is DNA. (Plomin, 2018: 11-12)
The problem with the mantra “nature and nurture” is not that it “runs the risk of sliding back into the mistaken view that the effects of genes and environment cannot be disentangled”—though that is one problem. The problem is how Plomin assumes how DNA works. That DNA can be disentangled from the environment presumes that DNA is environment-independent. But as Moore shows in his book The Dependent Gene—and as Schneider (2007) shows—“the very concept of a gene requires the environment“. Moore notes that “The common belief that genes contain context-independent “information”—and so are analogous to “blueprints” or “recipes”—is simply false” (quoted in Schneider, 2007). Moore showed in The Dependent Gene that twin studies are flawed, as have numerous other authors.
Lewkowicz (2012) argues that “genes are embedded within organisms which, in turn, are embedded in external environments. As a result, even though genes are a critical part of developmental systems, they are only one part of such systems where interactions occur at all levels of organization during both ontogeny and phylogeny.” Plomin—although he does not explicitly state it—is a genetic reductionist. This type of thinking can be traced back, most popularly, to Richard Dawkins’ 1976 book The Selfish Gene. The genetic reductionists can, and do, make the claim that organisms can be reduced to their genes, while developmental systems theorists claim that holism, and not reductionism, better explains organismal development.
The main thrust of Plomin’s Blueprint rests on (1) GWA studies and (2) PGSs/PRSs derived from the GWA studies. Ken Richardson (2017b) has shown that “some cryptic but functionally irrelevant genetic stratification in human populations, which, quite likely, will covary with social stratification or social class.” Richardson’s (2017b) argument is simple: Societies are genetically stratified; social stratification maintains genetic stratification; social stratification creates—and maintains—cognitive differentiation; “cognitive” tests reflect prior social stratification. This “cryptic but functionally irrelevant genetic stratification in human populations” is what GWA studies pick up. Richardson and Jones (2019) extend the argument and argue that spurious correlations can arise from genetic population structure that GWA studies cannot account for—even though GWA study authors claim that this population stratification is accounted for, social class is defined solely on the basis of SES (socioeconomic status) and therefore, does not capture all of what “social class” itself captures (Richardson, 2002: 298-299).
Plomin also heavily relies on the results of twin and adoption studies—a lot of it being his own work—to attempt to buttress his arguments. However, as Moore and Shenk (2016) show—and as I have summarized in Behavior Genetics and the Fallacy of Nature vs Nurture—heritability estimates for humans are highly flawed since there cannot be a fully controlled environment. Moore and Shenk (2016: 6) write:
Heritability statistics do remain useful in some limited circumstances, including selective breeding programs in which developmental environments can be strictly controlled. But in environments that are not controlled, these statistics do not tell us much. In light of this, numerous theorists have concluded that ‘the term “heritability,” which carries a strong conviction or connotation of something “[in]heritable” in the everyday sense, is no longer suitable for use in human genetics, and its use should be discontinued.’ 31 Reviewing the evidence, we come to the same conclusion.
Heritability estimates assume that nature (genes) can be separated from nurture (environment), but “the very concept of a gene requires the environment” (Schneider, 2007) so it seems that attempting to partition genetic and environmental causation of any trait T is a fool’s—reductionist—errand. If the concept of gene depends on and requires the environment, then how does it make any sense to attempt to partition one from the other if they need each other?
Let’s face it: Plomin, in this book Blueprint is speaking like a biological reductionist, though he may deny the claim. The claims from those who push PRS and how it can be used for precision medicine are unfounded, as there are numerous problems with the concept. Precision medicine and personalized medicine are similar concepts, though Joyner and Paneth (2015) are skeptical of its use and have seven questions for personalized medicine. Furthermore, Joyner, Boros and Fink (2018) argue that “redundant and degenerate mechanisms operating at the physiological level limit both the general utility of this assumption and the specific utility of the precision medicine narrative.” Joyner (2015: 5) also argues that “Neo-Darwinism has failed clinical medicine. By adopting a broader perspective, systems biology does not have to.”
Janssens and Joyner (2019) write that “Most [SNP] hits have no demonstrated mechanistic linkage to the biological property of interest.” Researchers can show correlations between disease phenotypes and genes, but they cannot show causation—which would be mechanistic relations between the proposed genes and the disease phenotype. Though, as Kampourakis (2017: 19), genes do not cause diseases on their own, they only contribute to its variation.
Edit: Take also this quote from Plomin and Stumm (2018) (quoted by Turkheimer):
GPS are unique predictors in the behavioural sciences. They are an exception to the rule that correlations do not imply causation in the sense that there can be no backward causation when GPS are correlated with traits. That is, nothing in our brains, behaviour or environment changes inherited differences in DNA sequence. A related advantage of GPS as predictors is that they are exceptionally stable throughout the life span because they index inherited differences in DNA sequence. Although mutations can accrue in the cells used to obtain DNA, like any cells in the body these mutations would not be expected to change systematically the thousands of inherited SNPs that contribute to a GPS.
Turkheimer goes on to say that this (false) assumption by Plomin and Stumm (2018) assumes that there is no top-down causation—i.e., that phenotypes don’t cause genes, or there is no causation from the top to the bottom. (See the special issue of Interface Focus for a slew of articles on top-down causation.) Downward causation exists in biological systems (Noble, 2012, 2017), as does top-down. The very claim that “nothing in our brains, behaviour or environment changes inherited differences in DNA sequence” is ridiculous! This is something that, of course, Plomin did not discuss in Blueprint. But in a book that, supposedly, shows “how DNA makes us who we are”, why not discuss epigenetics? Plomin is confused, because DNA methylation impacts behavior and behavior impacts DNA methylation (Lerner and Overton, 2017: 114). Lerner and Overtone (2017: 145) write that:
… it should no longer be possible for any scientist to undertake the procedure of splitting of nature and nurture and, through reductionist procedures, come to conclusions that the one or the other plays a more important role in behavior and development.
Plomin’s reductionist takes, therefore again, fail. Plomin’s “reluctance” to discuss “tangential topics” to “inherited DNA differences” included epigenetics (Plomin, 2018: 12). But it seems that his “reluctance” to discuss epigenetics was a downfall in his book as epigenetic mechanisms can and do make a difference to “inherited DNA differences” (see for example, Baedke, 2018, Above the Gene, Beyond Biology: Toward a Philosophy of Epigenetics and Meloni, 2019, Impressionable Biologies: From the Archaeology of Plasticity to the Sociology of Epigenetics see also Meloni, 2018). The genome can and does “react” to what occurs to the organism in the environment, so it is false that “nothing in our brains, behaviour or environment changes inherited differences in DNA sequence” (Plomin and Stumm, 2018), since our behavior and actions can and do methylate our DNA (Meloni, 2014) which falsifies Plomin’s claim and which is why he should have discussed epigenetics in Blueprint. End Edit
Conclusion
So the main premise of Plomin’s Blueprint is his two claims: (1) that DNA is a fortune teller and (2) that personal genomics is a fortune-telling device. He draws these big claims from PGS/PRS studies. However, over 80 percent of GWA studies have been done on European populations. And, knowing that we cannot use these datasets on other, non-European datasets, greatly hampers the uses of PGS/PRS in other populations—although the PGS/PRS are not that useful in and of itself for European populations. Plomin’s whole book is a reductionist screed—“Sure, other factors matter, but DNA matters more” is one of his main claims. Though, a priori, since there is no privileged level of causation, one cannot privilege DNA over any other developmental variables (Noble, 2012). To understand disease, we must understand the whole system and how when one part of the system becomes dysfunctional how it affects other parts of the system and how it runs. The PGS/PRS hunts are reductionist in nature, and the only answer to these reductionist paradigms are new paradigms from systems biology—one of holism.
Plomin’s assertions in his book are gleaned from highly confounded GWA studies. Plomin also assumes that we can disentangle nature and nurture—like all reductionists. Nature and nurture interact—without genes, there would be an environment, but without an environment, there would be no genes as gene expression is predicated on the environment and what occurs in it. So Plomin’s reductionist claim that “Our future is DNA” is false—our future is studying the interactive developmental system, not reducing it to a sum of its parts. Holistic biology—systems biology—beats reductionist biology—the Neo-Darwinian Modern Synthesis.
DNA is not a blueprint nor is it a fortune teller and personal genomics is not a fortune-telling device. The claim that DNA is a blueprint/fortune teller and personal genomics is a fortune-telling device come from Plomin and are derived from highly flawed GWA studies and, further, PGS/PRS. Therefore Plomin’s claim that DNA is a blueprint/fortune teller and personal genomics is a fortune-telling device are false.
(Also read Erick Turkheimer’s 2019 review of Plomin’s book The Social Science Blues, along with Steve Pitteli’s review Biogenetic Overreach for an overview and critiques of Plomin’s ideas. And read Ken Richardson’s article It’s the End of the Gene As We Know It for a critique of the concept of the gene.)
Gene-Selectionism vs. Developmental Systems Theory
2300 words
Two dominant theories exist in regard to development, the “gene’s eye view—gene selectionism (GS)—and the developmental view—developmental systems theory (DST). GS proposes that there are two fundamental processes in regard to evolution: replication and interaction. Replicators (the term was coined by Dawkins) are anything that is copied into the next generation whereas interactors (vehicles) are things that only exist to ensure the replicators’ survival. Thus, Dawkins (1976) proposes a distinction between the “vehicle” (organism) and its “riders/replicators” (the genes).
Gene selectionism
Gene selectionists propose a simple hypothesis: evolution through the differential survival of genes, its main premise being that the “gene” is “the ultimate, fundamental unit of natural selection.” Dusek (1999: 156) writes that “Gene selection claims that genes, not organisms, groups of organisms or species, are selected. The gene is considered to be the unit of selection.” The view of gene selectionists is best—and most popularly put—by Richard Dawkins’ seminal book The Selfish Gene (1976), in which he posits that genes “compete” with each other, and that our “selfish actions” are the result of our genes attempting to replicate to the next generation, relegating our bodies to disposable “vehicles” that only house the “replicators” (or “drivers).
Though, just because one is a gene selectionist does not necessarily mean that they are a genetic determinist (both views will be argued below). Gene selectionists are comitted to the view that genes make a distinctive contribution toward building interactors. Dawkins (1982) claims that genetic determinism is not a problem in regard to gene selectionism. Replicators (genes) have a special status to gene selectionists. Gene selectionists argue that adaptive evolution only occurs through cumulative selection, while only the replicators persist through the generations. Gene selectionists do not see organisms as replicators since genes—and not organisms—are what is replicated according to the view.
The gene selectionist view (Dawkins’ 1976 view) can also be said to apply what Okasha (2018) terms “agential thinking”. “Agential thinking” is “treating an evolved organism as if it were an agent pursuing a goal, such as survival or reproduction, and treating its phenotypic traits, including its behaviour, as strategies for achieving that goal, or furthering its biological interests” (Okasha, 2018: 12). Dawkins—and other gene selectionists—treat genes as if they have agency, speaking of “intra-genomic conflict”, as if genes are competing with each other (sure, it’s “just a metaphor”, see below).
Okasha (2018: 71) writes:
To see how this distinction relates to agential thinking, note that every gene is necessarily playing a zero-sum game against other alleles at the same locus: it can only spread in the population if they decline. Therefore every gene, including outlaws, can be thought of as ‘trying’ to outcompete alleles, or having this as its ultimate goal.
Selfish genes also have intermediate goals, which are to maximize fitness, which is done through expression in the organismic phenotype.
Thus, according to Okasha (2018: 73), “… selfish genetic elements have phenotypic effects which can be regarded as adaptations, but only if we apply the notions of agent, benefit, and goal to genes themselves”, though “… only in an evolutionary context [does] it [make] sense to treat genes as agent-like and credit them with goals and interests.” It does not “make sense to treat genes as even “agent-like and credit them with goals and interests since they can only be attributed to humans.
Other genes have as their intermediate goal to enhance the fitness of their host organism’s relatives, by causing altruistic behaviour [genes can’t cause altruistic behavior; it is an action]. However, a small handful of genes have a different intermediate goal, namely to increase their own transmission in their host organism’s gametes, for example, by biasing segregation in their favour, or distorting the sex-ratio, or transposing to new sites in the genome. These are outlaws, or selfish genetic elements.If oulaws are absent or are effectively suppressed, then the genes within a single organism have a common (intermediate) goal, so will cooperate: each gene can onluy benefit by itself by benefiting the whole organism. Agential thinking then can be applied to the organism itself. The organism’s goal—maximizing its fitness—then equates to the intermediate goal of each of the genes within it. (Okasha, 2018: 72)
Attributing agential thinking to anything other than humans is erroneous, since genes are not “selfish.”
The selfish gene is one of the main theories that define the neo-Darwinian paradigm and it is flat out wrong. Genes are not ultimate causes, as the crafters of the neo-Darwinian Modern Synthesis (MS) propose, genes are resources in a dynamic system and can thusly only be seen as causes in a passive, not active, sense (Noble, 2011).
Developmental systems
The alternative to the gene-centric view of evolution is that of developmental systems theory (DST), first proposed by Oyama (1985).
The argument for DST is simple:
(1) Organisms obviously inherit more than DNA from their parents. Since organisms can behave in ways that alter the environment, environments are also passed onto offspring. Thus, it can be said that genes are not the only things inherited, but a whole developmental matrix is.
(2) Genes, according to the orthodox view of the MS, interact with many other factors for development to occur, and so genes are not the only thing that help ‘build’ the organism. Genes can still play some “privileged” role in development, in that they “control”, “direct” or “organize” everything else, but this is up to gene-selectionists to prove. (See Noble, 2012.)
(3) The common claim that genes contain “information” (that is, context-independent information) is untenable, since every reconstruction of genes contain development about information applies directly to all other developmental outcomes. Genes cannot be singled out as privileged causes in development.
(4) Other attempts—such as genes are copied more “directly—are mistaken, since they draw a distinction between development and other factors but fail.
(5) Genes, then, cannot be privileged in development, and are no different than any other developmental factor. Genes, in fact, are just passive templates for protein construction, waiting to be used by the system in a context-dependent fashion (see Moore, 2002; Schneider, 2007). The entire developmental system reconstructs itself “through numerous independent causal pathways” (Sterelny and Griffiths, 1999: 109).
DNA is not the only thing inherited, and the so-called “famed immortality of DNA is actually a property of cells [since] [o]nly cells have the machinery to correct frequent faults that occur in DNA replication.” The thing about replication, though, is that “DNA and the cell must replicate together” (Noble, 2017: 238). A whole slew of developmental tools are inherited and that is what constructs the organism; organisms are, quite obviously, constructed not by genes alone.
Developmental systems, as described by Oyama (1985: 49) do not “have a final form, encoded before its starting point and realized at maturity. It has, if one focuses finely enough, as many forms as time has segments.” Oyama (1985: 61) further writes that “The function of the gene or any other influence can be understood only in relation to the system in which they are involved. The biological relevance or any influence, and therefore the very “information” it conveys, is jointly determined, frequently in a statistically interactive, not additive, manner, by that influence and the system state it influences.”
DNA is, of course, important. For without it, there would be nothing for the cell to read (recall how the genome is an organ of the cell) and so no development would occur. DNA is only “information” about an organism only in the process of cellular functioning.
The simple fact of the matter is this: the development of organs and tissues are not directly “controlled” by genes, but by the exchange signals of the cells. “Details notwithstanding, what is important to note is that whatever kinds of signals it sends out depends on the kind of signals it receives from its immediate environment. Therefore, neighboring cells are interdependent, and its local interactions among cells that drive the developmental processes” (Kampourakis, 2017: 173).
The fact of the matter is that whether or not a trait is realized depends on the developmental processes (and the physiologic system itself) and the environment. Kampourakis, just like Noble (2006, 2012, 2017) pushes a holistic view of development and the system. Kampourakis (2017: 184) writes:
What genetics research consistently shows is that biological phenomena should be approached holistically. at various levels. For example, as genes are expressed and produce proteins, and some of these proteins regulate or affect gene expression, there is absolutely no reason to privilege genes over proteins. This is why it is important to consider developmental processes in order to undertand how characters and disease arise. Genes cannot be considered alone but only in the broader context (cellular, organismal, environmental) in which they exist. And both characters and disease in fact develop; they are not just produced. Therefore, reductionism, the idea that genes provide the ultimate explanation for characters and disease, is also wrong. In order to understand such phenomena, we need to consider influence at various levels of organization, both bottom-up and top-down. This is why current research has adopted a systems biology approach (see Noble, 2006; Voit, 2016 for accessible introductions).
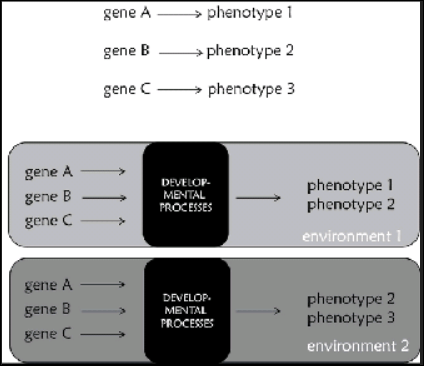
All this shows that developmental processes and interactions play a major role in shaping characters. Organisms can respond to changing environments through changes in their development and eventually their phenotypes. Most interestingly, plastic responses of this kind can become stable and inherited by their offspring. Therefore, genes do not predetermine phenotypes; genes are implicated in the development of phenotypes only through their products, which depends on what else is going on within and outside cells (Jablonka, 2013). It is therefore necessary to replacr the common representation of gene function presented in Figure 9.6a, which we usually find in the public sphere, with others that consider development, such as the one in figure 9.6b. Genes do not determine characters, but they are implicated in their development. Genes are resources that provide cells with a generative plan about the development of the organism, and have a major role in this process through their products. This plan is the resouce for the production of robust developmental outcomes that are at the same time plastic enough to accomodate changes stemming from environmental signals.
Figure 9.6 (a) The common representation of gene function: a single gene determines a single phenotype. It should be clear by what has been present in the book so far that is not accurate. (b) A more accurate representation of gene function that takes development and environment into account. In this case, a phenotype is propduced in a particular environment by developmental processes in which genes are implicated. In a different environment the same genes might contribute tothe development of a different phenotype. Note the “black box” of development.
[Kampourakis also writes on page 188, note 3]
In the original analogy, Wolpert (2011, p. 11) actually uses the term “program.” However, I consider the term “plan” as more accurate and thus more appropriate. In my view, the term “program” impies instructions and their implimentation, whereas the term “plan” is about instructions only. The notion of a genetic program can be very misleading because it implies that, if it were technically feasible, it would be possible to compute an organism by reading the DNA sequence alone (see Lewontin, 2000, pp. 17-18).
Kampourakis is obviously speaking of a “plan” in a context-dependent manner since that is the only way that genes/DNA contain “information” (Moore, 2002; Schneider, 2007). The whole point is that genes, to use Noble’s terminology, are “slaves” to the system, since they are used by and for the (physiological) system. Developmental systems theory is a “wholeheartedly epigenetic approach to development, inheritance and evolution” (Hochman and Griffiths, 2015).
This point is driven home by Richardson (2017:111):
And how did genes eventually become established? Probably not at all as the original recipes, designers, and controllers of life. Instead they arose as templates for molecular components used repeatedly in the life of the cell and the organism: a kind of facility for just-in-time production of parts needed on a recurring basis. Over time, of course, the role of these parts themselves evolved to become key players in the metabolism of the call—but as part of a team, not the boss.
[…]
It is not surprising, then, that we find that variation in form and function has, for most traits, only a tenuous relationship with variation in genes.
[And also writes on page 133]:
There is no direct command line between environments and genes or between genes and phenotypes. Predictions and decisions about form and variation are made through a highly evolved dynamical system. That is why ostensibly the same environment, such as hormonal signal, can initiate a variaety of responses like growth, cell division, differentiation, and migration, depending on deeper context. This reflects more than fixes responses from fixed information in genes, something fatally overlooked in the nature-nurture debate
(Also read Richardson’s article So what is a gene?)
Conclusion
The gene-selectionist point-of-view entails too many (false) assumptions. The DST point of view, on the other hand, does not fall prey to the pitfalls of the gene-selectionist POV; Developmental systems theorists look at the gene, not as the ultimate causes of development—and, along with that, only changes in gene frequency driving evolutionary change—but only as products to be used by and for the system. Genes can only be looked at in terms of development, and in no other way (Kamporuakis, 2017; Noble, 2017). Thus, the gene-selectionists are wrong; the main tenet of the neo-Darwinian Modern Synthesis, gene-selectionism—the selfish gene—has been refuted (Jablonka and Lamb, 2005; Noble, 2006, 2011). The main tenets of the neo-Darwinian Modern Synthesis have been refuted, and so it is now time to replace the Modern Synthesis with a new view of evolution: one that includes the role of genes and development and the role of epigenetics on the developmental system. The gene-selectionist view champions an untenable view of the gene: that the gene is priviliged above any other developmental variables, but Noble and Kampourakis show that this is not the case, since DNA is inherited with the cell; the cell is what is “immortal” to use the language of Dawkins—not DNA itself.
A priori, there is no privileged level of causation, and this includes the gene, which so many place at the top of the hierarchy (Noble, 2012).
Do Genes and Polymorphisms Explain the Differences between Eastern and Western Societies?
2150 words
In 2012, biologist Hippokratis Kiaris published a book titled Genes, Polymorphisms, and the Making of Societies: How Genetic Behavioral Traits Influence Human Cultures. His main point is that “the presence of different genes in the corresponding people has actually dictated the acquisition of these distinct cultural and historical lines, and that an alternative outcome might be unlikely” (Kiaris, 2012: 9). This is a book that I have not seen discussed in any HBD blog, and based on the premise of the book (how it purports to explain behavioral/societal outcomes between Eastern and Western society) you would think it would be. The book is short, and he speaks with a lot of determinist language. (It’s worth noting he does not discuss IQ at all.)
In the book, he discusses how genes “affect” and “dictate” behavior which then affects “collective decisions and actions” while also stating that it is “conceivable” that history, and what affects human decision-making and reactions, are also “affected by the genetic identity of the people involved” (Kiaris, 2012: 11). Kiaris argues that genetic differences between Easterners and Westerners are driven by “specific environmental conditions that apparently drove the selection of specific alleles in certain populations, which in turn developed particular cultural attitudes and norms” (Kiaris, 2012: 91).
Kiaris attempts to explain the societal differences between the peoples who adopted Platonic thought and those who adopted Confucian thought. He argues that differences between Eastern and Western societies “are not random and stochastic” but are “dictated—or if this is too strong an argument, they are influenced considerably—by the genes that these people carry.” So, Kiaris says, “what we view as a choice is rather the complex and collective outcome of the influence of people’s specific genes combined with the effects of their specific environment … [which] makes the probability for rendering a certain choice distinct between different populations” (Kiaris, 2012: 50).
The first thing that Kiaris discusses (behavior wise) is DRD4. This allele has been associated with miles migrated from Africa (with a correlation of .85) along with novelty-seeking and hyperactivity (which may cause the association found with DRD4 frequency and miles migrated from Africa (Chen et al, 1999). Kiaris notes, of course, that the DRD4 alleles are unevenly distributed across the globe, with people who have migrated further from Africa having a higher frequency of these alleles. Europeans were more likely to have the “novelty-seeking” DRD7 compared to Asian populations (Chang et al, 1996). But, Kiaris (2012: 68) wisely writes (emphasis mine):
Whether these differences [in DRD alleles] represent the collective and cumulative result of selective pressure or they are due to founder effects related to the genetic composition of the early populations that inhabited the corresponding areas remains elusive and is actually impossible to prove or disprove with certainty.
Kiaris then discusses differences between Eastern and Western societies and how we might understand these differences between societies as regards novelty-seeking and the DRD4-7 distribution across the globe. Westerners are more individualistic and this concept of individuality is actually a cornerstone of Western civilization. The “increased excitability and attraction to extravagance” of Westerners, according to Kiaris, is linked to this novelty-seeking behavior which is also related to individualism “and the tendency to constantly seek for means to obtain satisfaction” (Kiaris, 2012: 68). We know that Westerners do not shy away from exploration; after all, the West discovered the East and not vice versa.
Easterners, on the other hand, are more passive and have “an attitude that reflects a certain degree of stoicism and makes life within larger—and likely collectivistic—groups of people more convenient“. Easterners, compared to Westerners, take things “the way they are” which “probably reflects their belief that there is not much one can or should do to change them. This is probably the reason that these people appear rigid against life and loyal, a fact that is also reflected historically in their relatively high political stability” (Kiaris, 2012: 68-69).
Kiaris describes DRD4 as a “prototype Westerner’s gene” (pg 83), stating that the 7R allele of this gene is found more frequently in Europeans compares to Asians. The gene has been associated with increased novelty-seeking, exploratory activity and human migrations, along with liberal ideology. These, of course, are cornerstones of Western civilization and thought, and so, Kiaris argues that the higher frequency of this allele in Europeans—in part—explains certain societal differences between the East and West. Kiaris (2012: 83) then makes a bold claim:
All these features [novelty-seeking, exploratory activity and migration] indeed tend to characterize Westerners and the culutral norms they developed, posing the intriguing possibility that DRD4 can actually represent a single gene that can “predispose” for what we understand as the stereotypic Western-type behavior. Thus, we could imagine that an individual beating the 7-repeat allele functions more efficiently in Western society while the one without this allele would probably be better suited to a society with Eastern-like structure. Alternatively, we could propose that a society with more individuals bearing the 7-repeat allele is more likely to have followed historical lines and choices more typical of a Western society, while a population with a lower number (or deficient as it is the actual case with Easterners) of individuals with the 7-repeat allele would more likely attend to the collective historical outcome of Eaasterners.
Kiaris (2012: 84) is, importantly, skeptical that having a high number of “novelty-seekers” and “explorers” would lead to higher scientific achievement. This is because “attempts to extrapolate from individual characteristics to those of a group of people and societies possess certain dangers and conceptual limitations.”
Kiaris (2012: 86) says that “collectivistic behavior … is related to the activity of serotonin.” He then goes on to cite a few instances of other polymorphisms which are associated with collective behavior as well. Goldman et al (2010) show ethnic differences in the l and s alleles (from Kiaris, 2012: 86):
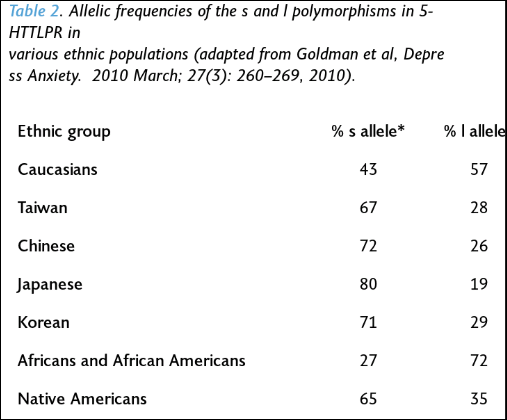
It should also be noted that populations (Easterners) that had a higher frequency of the s allele had a lower prevalence of depression than Westerners. So Western societies are more likely to “suffer more frequently from various manifestations of depression and general mood disorders than those of Eastern cultures (Chiao & Blizinsky, 2010)” (Kiaris, 2012: 89).
As can be seen from the table above, Westerners are more likely to have the l allele than Easterners, which should subsequently predict higher levels of happiness in Western compared to Eastern populations. However, “happiness” is, in many ways, subjective; so how would one find an objective way to measure “happiness” cross-culturally? However, Kiaris (2012: 94) writes: “Intuitively speaking, though, I have to admit that I would rather expect Asians to be happier, in general, than Westerners. I cannot support this by specific arguments, but I think the reason for that is related to the individualistic approach of life that the people possess in Western societies: By operating under individualistic norms, it is unavoidably stressful, a condition that operates at the expense of the perception of individuals’ happiness.”
Kiaris discusses catechol-O-methyltransferase (COMT), which is an enzyme responsible for the inactivation of catecholamines. Catecholamines are the hormones dopamine, adrenaline, and noradrenaline. These hormones regulate the “fight or flight” function (Goldstein, 2011). So since catecholamines play a regulatory role in the “fight or flight” mechanism, increased COMT activity results in lower dopamine levels, which is then associated with better performance.
“Warriors” and “worriers” are intrinsically linked to the “fight or flight” mechanism. A “warrior” is someone who performs better under stress, achieves maximal performance despite threat and pain, and is more likely to act efficiently in a threatening environment. A “worrier” is “someone that has an advantage in memory and attention tasks, is more exploratory and efficient in complex environments, but who exhibits worse performance under stressful conditions (Stein et al., 2006)” (Kiaris, 2012: 102).
Kiaris (2012: 107) states that “at the level of society, it can be argued that the specific Met-bearing COMT allele contributes to the buildup of Western individualism. Opposed to this, Easterners’ increased frequency of the Val-bearing “altruistic” allele fits quite well with the construction of a collectivistic society: You have to be an altruist at some degree in order to understand the benefits of collectivism. By being a pure individualist, you only understand “good” as defined and reflected by your sole existence.”
So, Kiaris’ whole point is thus: there are differences in polymorphic genes between Easterners and Westerners (and are unevenly distributed) and that differences in these polymorphisms (DRD4, HTT, MAOA, and COMT) explain behavioral differences between behaviors in Eastern and Western societies. So the genetic polymorphisms associated with “Western behavior” (DRD4) are associated with increased novelty-seeking, tendency for financial risk-taking, distance of OoA migration, and liberal ideology. Numerous different MAOA and 5-HTT polymorphisms are associated with collectivism (e.g., Way and Lieberman, 2006 for MAOA and collectivism). The polymorphism in COMT more likely to be found in Westerners predisposes for “worrier’s behavior”. Furthermore, certain polymorphisms of the CHRNB3 gene are more common in all of the populations that migrated out of Africa, which predisposed for leaders—and not follower—behavior.
| Trait | Gene | Allele frequency |
| Novelty seeking | DRD4 | 7-repeat novelty seeking allele more common in the West |
| Migration | DRD4 | 7-repeat allele is associated with distance from Africa migration |
| Nomads/settlers | DRD4 | 7-repeat allele is associated with nomadic life |
| Political ideology | DRD4 | 7-repeat allele is more common in liberals |
| Financial risk taking | DRD4 | 7-repeat allele is more common in risk takers |
| Individualism/Collectivism | HTT | s allele (collectivistic) of 5-HTT is more common in the East |
| Happiness | HTT | l allele has higher prevalence in individuals happy with their life |
| Individualism/Collectivism | MAOA | 3-repeat allele (collectivistic) more common in the East) |
| Warrior/Worrier | COMT | A-allele (worrier) more common in the West |
| Altruism | COMT | G-allele (warrior) associated with altruism |
| Leader/Follower | CHRBN3 | A-allele (leader) more common in populations Out-of-Africa |
The table above is from Kiaris (2012: 117) who lays out the genes/polymorphisms discussed in his book—what supposedly shows how and why Eastern and Western societies are so different.
Kiaris (2012: 141) then makes a bold claim: “Since we know now that at least a fraction (and likely more than that) of our behavior is due to our genes“, actually “we” don’t “know” this “now”.
The takeaways from the book are: (1) populations differ genetically; (2) since populations differ genetically, then genetic differences correlated with behavior should show frequency differences between populations; (3) since these populations show both behavioral/societal differences and they also differ in genetic polymorphisms which are then associated with that behavior, then those polymorphisms are, in part, a cause of that society and the behavior found in it; (4) therefore, differences in Eastern and Western societies are explained by (some) of these polymorphisms discussed.
Now for a simple rebuttal of the book:
“B iff G” (behavior B is possible if and only if a specific genotype G is instantiated) or “if G, then necessarily B” (genotype G is a sufficient cause for behavior B). Both claims are false; genes are neither a sufficient or necessary cause for any behavior. Genes are, of course, a necessary pre-condition for behavior, but they are not needed for a specific behavior to be instantiated; genes can be said to be difference makers (Sterelny and Kitcher, 1988) (but see Godfrey-Smith and Lewontin, 1993 for a response). These claims cannot be substantiated; therefore, the claims that “if G, then necessarily B” and “B iff G” are false, it cannot be shown that genes are difference makers in regard to behavior, nor can it be shown that particular genes or whatnot.
Conclusion
I’m surprised that I have not come across a book like this sooner; you would expect that there would be a lot more written on this. This book is short, it discusses some good resources, but the conclusions that Kiaris draws, in my opinion, will not come to pass because genes are not neccesary nor sufficient cause of any type of behavior, nor can it be shown that genes are causes of any behavior B. Behavioral differences between Eastern and Western societies, logically, cannot come down to differences in genes, since they are neither necessary nor sufficient causes of behavior (genes are neccessary pre-conditions for behavior, since without genes there is no organism, but genes cannot explain behavior).
Kiaris attempts to show how and why Eastern and Western societies became so different, how and why Western societies are dominated by “Aristotle’s reason and logic”, while Eastern lines of thought “has been dominated by Confucious’s harmony, collectivism, and context dependency” (Kiaris, 2012: 9). While the book is well-written and researched (he talks about nothing new if you’re familiar with the literature), Kiaris fails to prove his ultimate point: that differences in genetic polymorphisms between individuals in different societies explain how and why the societies in question are so different. Though, it is not logically possible for genes to be a necessary nor sufficient cause for any behavior. Kiaris talks like a determinist, since he says that “the presence of different genes in the corresponding people has actually dictated the acquisition of these distinct cultural and historical lines, and that an alternative outcome might be unlikely” (Kiaris, 2012: 9), though that is just wishful thinking: if we were able to start history over again, things would occur differently, “the presence of different genes in the corresponding people” be dammed, since genes do not cause behavior.
DNA is not a “Blueprint”
2200 words
Leading behavior geneticist Robert Plomin is publishing “Blueprint: How DNA Makes Us Who We Are” in October of 2018. I, of course, have not read the book yet. But if the main thesis of the book is that DNA is a “code”, “recipe”, or “blueprint”, then that is already wrong. This is because presuming that DNA is any of the three aforementioned things marries one to certain ideas, even if they themselves do not explicitly state them. Nevertheless, Robert Plomin is what one would term a “hereditarian”, meaning that he believes that genes—more than environment—shape an individual’s psychological and other traits. (That’s a false dichotomy, though.) In the preview for the book at MIT Press, they write:
In Blueprint, behavioral geneticist Robert Plomin describes how the DNA revolution has made DNA personal by giving us the power to predict our psychological strengths and weaknesses from birth. A century of genetic research shows that DNA differences inherited from our parents are the consistent life-long sources of our psychological individuality—the blueprint that makes us who we are. This, says Plomin, is a game-changer. It calls for a radical rethinking of what makes us who were are.
Genetics accounts for fifty percent of psychological differences—not just mental health and school achievement, but all psychological traits, from personality to intellectual abilities. Nature defeats nurture by a landslide.
Plomin explores the implications of this, drawing some provocative conclusions—among them that parenting styles don’t really affect children’s outcomes once genetics is taken into effect. Neither tiger mothers nor attachment parenting affects children’s ability to get into Harvard. After describing why DNA matters, Plomin explains what DNA does, offering readers a unique insider’s view of the exciting synergies that came from combining genetics and psychology.
I won’t get into most of these things today (I will wait until I read the book for that), but this will be just an article showing that DNA is, in fact, not a blueprint, and DNA is not a “code” or “recipe” for the organism.
It’s funny that the little blurb says that “Nature defeats nurture by a landslide“, because, as I have argued at length, nature vs nurture is a false dichotomy (See Oyama, 1985, 2000, 1999; Moore, 2002; Schneider, 2007; Moore, 2017). Nature vs nurture is the battleground that the false dichotomy of genes vs environment is fought on. However, it makes no sense to partition heritability estimates if it is indeed true that genes interact with environment—that is, if nature interacts with nurture.
DNA is also called “the book of life”. For example, in her book The Epigenetics Revolution: How Modern Biology Is Rewriting Our Understanding of Genetics, Disease, and Inheritance, Nessa Carey writes that “There’s no debate that the DNA blueprint is a starting point” (pg 16). This, though, can be contested. “But the promise of a peep into the ‘book of life’ leading to a cure for all diseases was a mistake” (Noble, 2017: 161).
Developmental psychologist and cognitive scientist David S. Moore concurs. In his book The Developing Genome: An Introduction to Behavioral Epigenetics, he writes (pg 45):
So, although I will talk about genes repeatedly in this book, it is only because there is no other convenient way to communicate about contemporary ideas in molecular biology. And when I refer to gebe, I will be talking about a segment or segments of DNA containing sequence information that is used to help construct a protein (or some other product that performs a biological function). But it is worth remembering that contemporary biologists do not mean any one thing when they talk about “genes”; the gene remains a fundementally hypothetical concept to this day. The common belief that there are things inside of us that constitute a set of instructions for building bodies and minds—things that are analogous to “blueprings” or “recipes”—is undoubedtly false. Instead, DNA segements often contain information that is ambiguous, and that must be edited or arranged in context-dependent ways before it can be used.
Still, other may use terms like “genes for” trait T. This, too, is incorrect. In his outstanding book Making Sense of Genes, Kostas Kamporakis writes (pg 19):
I also explain why the notion of “genes for,” in the vernacular sense, is not only misleading but also entirely inaccurate and scientifcally illegitamate.
[…]
First, I show that genes “operate” in the context of development only. This means that genes are impllicated in the development of characters but do not determine them. Second, I explain why single genes do not alone produce characters or disease but contribute to their variation. This means that genes can account for variation in characters but cannot alone explain their origin. Third, I show that genes are not the masters of the game but are subject to complex regulatory processes.
Genes can only be seen as passive templates, not ultimate causes (Noble, 2011), and they cannot explain the origin of different characters but can account for variation in physical characters. Genes only “do” something in the context of development; they are inert molecules and thusly cannot “cause” anything on their own.
Genes are not ‘for’ traits, but they are difference-makers for traits. Sterelny and Griffiths (1999: 102), in their book Sex and Death: An Introduction to Philosophy of Biology write:
Sterelny and Griffiths (1988) responded to the idea that genes are invisible to selection by treating genes as difference makers, and as visible to selection by virtue of the differences they make. In doing so, they provided a formal reconstruction of the “gene for” locution. The details are complex, but the basic intent of the reconstruction is simple. A certain allele in humans is an “allele for brown eyes” because, in standard environments, having that allele rather than alternatives typically available in the population means that your eyes will be brown rather than blue. This is the concpet of a gene as a difference maker. It is very important to note, however, that genes are context-sensitive difference makers. Their effects depend on the genetic, cellular, and other features of their environment.
(Genes can be difference makers for physical traits, but not for psychological traits because no psychophysical laws exist, but I’ll get to that in the future.)
Note how the terms “context-sensitive” and “context-dependent” continue to appear. The DNA-as-blueprint statement presumes that DNA is context-independent, but we cannot divorce genes—whatever they are—from their context, since genes and environment, nature and nurture, are intertwined. (And it is even questioned if ‘genes’ are truly units of inheritance, see Fogle, 1990. Fogle, 2000 also argues to dispense with the concept of “gene” and that biologists should be using terms like intron, promoter region, and exon. Nevertheless, there is a huge disconnect with the term “gene” in molecular biology and classical genetics. Keller 2000 argues that there are still uses for the term “gene” and that we should not dispense with the term. I believe we should dispense with it.)
Susan Oyama (2000: 77) writes in her book The Ontogeny of Information:
“Though a plan implies action, it does not itself act, so if the genes are a blueprint, something else is the constructor-construction worker. Though blueprints are usually contrasted with building materials, the genes are quite easily conceptualized as templates for building tools and materials; once so utilized, of course, they enter the developmental process and influence its course. The point of the blueprint analogy, though, does not seem to be to illuminate developmental processes, but rather to assume them and, in celebrating their regularity, to impute cognitive functions to genes. How these functions are exercised is left unclear in this type of metaphor, except that the genetic plan is seen in some peculiar way to carry itself out, generating all the necessary steps in the necessary sequence. No light is shed on multiple developmental possibilities, species-typical or atypical.“
The Modern Synthesis is one of the causes for the genes-as-blueprints thinking; the Modern Synthesis has causation in biology wrong. Genes are not active causes, but they are passive templates, as argued by many authors. They, thus, cannot “cause” anything on their own.
In his 2017 book Dance to the Tune of Life: Biological Relativity, Denis Noble writes (pg 157):
As we saw earlier in this chapter, these triplet sequences are formed from any combination of the four bases U, C, A and G in RNA and T, C, A and G in DNA. They are often described as a genetic ‘code’, but it is important to understand that this usage of the word ‘code’ carries overtones that can be confusing.
A code was originally an intentional encryption used by humans to communicate. The genetic ‘code’ is not intentional in that sense. The word ‘code’ has unfortunately reinforced the idea that genes are active and even complete causes, in much the same was as a computer is caused to follow the instructions of a computer program. The more nuetral word ‘template’ would be better. Templates are used only when required (activated); they are not themselves active causes. The active causes lie within the cells themselves since they determine the expression patterns for the different cell types and states. These patterns are comminicated to the DNA by transcrption factors, by methylation patterns and by binding to the tails of histones, all of which influence the pattern and speed of transcription of different parts of the genome. If the word ‘instruction’ is useful here at all, it is rather that the cell instructs the genome. As Barbara McClintock wrote in 1984 after receiving her Nobel Prize, the genome is an ‘organ of the cell’, not the other way around.
Realising that DNA is under the control of the system has been reinforced by the discovery that cells use different start, stop and splice sites for producing different messenger RNAs from a single DNA sequence. This enables the same sequence to code different proteins in different cell types and under different conditions [here’s where context-dependency comes into play again].
Representing the direction of causality in biology the wrong way round is therefore confusing and has far-reaching conseqeunces. The causality is circular, acting both ways: passive causality by DNA sequences acting as otherwise inert templates, and active causality by the functional networks of interactions that determine how the genome is activated.
This takes care of the idea that DNA is a ‘code’. But what about DNA being a ‘blueprint’, that all of the information is contained in the DNA of the organism before conception? DNA is clearly not a ‘program’, in the sense that all of the information to construct the organism exists already in DNA. The complete cell is also needed, and its “complex structures are inherited by self-templating” (Noble, 2017: 161). Thus, the “blueprint” is the whole cell, not just the genome itself (remember that the genome is an organ of the cell).
Lastly, GWA studies have been all the rage recently. However, there is only so much we can learn just from association studies, before we need to turn to the physiological sciences for functional analyses. Indeed, Denis Noble (2018) writes in a new editorial:
As with the results of GWAS (genome-wide association studies) generally, the associations at the genome sequence level are remarkably weak and, with the exception of certain rare genetic diseases, may even be meaningless (13, 21). The reason is that if you gather a sufficiently large data set, it is a mathematical necessity that you will find correlations, even if the data set was generated randomly so that the correlations must be spurious. The bigger the data set, the more spurious correlations will be found (3).
[…]
The results of GWAS do not reveal the secrets of life, nor have they delivered the many cures for complex diseases that society badly needs. The reason is that association studies do not reveal biological mechanisms. Physiology does. Worse still, “the more data, the more arbitrary, meaningless and useless (for future action) correlations will be found in them” is a necessary mathematical statement (3).
Nor does applying a highly restricted DNA sequence-based interpretation of evolutionary biology, and its latest manifestation in GWAS, to the social sciences augur well for society.
It is further worth noting that there is no privileged level of causation in biological systems (Noble, 2012)—a priori, there is no justification to privilege one system over another in regard to causation, so saying that one level of the organism is “higher” than another (for instance, saying that genes are, and should be, privileged over the environment or any other system in the organism regarding causation) is clearly false, since there is upwards and downwards causation, influencing all levels of the system.
In sum, it is highly misleading to refer to DNA as “blueprints”, a “code”, or a “recipe.” Referring to DNA in this way means that one presumes that DNA can be divorced from its context—that it does not work together with the environment. As I have argued in the past, association studies will not elucidate genetic mechanisms, nor will heritability estimates (Richardson, 2012). We need physiological testing for these functional analyses, and association studies like GWAS and even heritability estimates don’t tell us this type of information (Panofsky, 2014). So, it seems, that what Plomin et al are looking for that they assume are “in the genes”, are not there, because they use a false model of the gene (Burt, 2015; Richardson, 2017). Genes are resources—templates to be used by and for the system—not causes of traits and development. They can account for differences in variation, but cannot be said to be the origin of trait differences. Genes can be said to be difference makers, but knowing whether or not they are difference makers for behavior, in my opinion, cannot be known.
(For further information on genes and what they do, reach Chapters Four and Five of Ken Richardson’s book Genes, Brains, and Human Potential: The Science and Ideology of Intelligence. Plomin himself seems to be a reductionist, and Richardson took care of that paradigm in his book. Lickliter (2018) has a good review of the book, along with critiques of the reductionist paradigm that Plomin et al follow.)
Genotypes, Athletic Performance, and Race
2050 words
Everyone wants to know the keys to athletic success, however, as I have argued in the past, to understand elite athletic performance, we must understand how the system works in concert with everything—especially in the environments the biological system finds itself in. To reduce factors down to genes, or training, or X or Y does not make sense; to look at what makes an elite athlete, the method of reductionism, while it does allow us to identify certain differences between athletes, it does not allow us to appreciate the full-range of how and why elite athletes differ in their sport of choice. One large meta-analysis has been done on the effects of a few genotypes on elite athletic performance, and it shows us what we already know (blacks are more likely to have the genotype associated with power performance—so why are there no black Strongmen or any competitors in the World’s Strongest Man?). A few studies and one meta-analysis exist, attempting to get to the bottom of the genetics of elite athletic performance and, while it of course plays a factor, as I have argued in the past, we must take a systems view of the matter.
One 2013 study found that a functional polymorphism in the angiotensinogen (ATG) region was 2 to 3 times more common in elite power athletes than in (non-athlete) controls and elite endurance athletes (Zarebska et al, 2013). This sample tested was Polish, n = 223, 156 males, 67 females, and then they further broke down their athletic sample into tiers. They tested 100 power athletes (29 100-400 m runners; 22 powerlifters; 20 weightlifters; 14 throwers and 15 jumpers) and 123 endurance athletes (4 tri-athletes; 6 race walkers; 14 road cyclists; 6 15 to 50 m cross-country skiers; 12 marathon runners; 53 rowers; 17 3 to 10 km runners; and 11 800 to 1500 m swimmers).
Zarebska et al (2013) attempted to replicate previous associations found in other studies (Buxens et al, 2009) most notably the association with the M235T polymorphism in the AGT (angiotensinogen) gene. Zarebska et al’s (2013) main finding was that there was a higher representation of elite power athletes with the CC and C alleles of the M235T polymorphism compared with endurance athletes and controls, which suggests that the C allele of the M235T gene “may be associated with a predisposition to power-oriented
events” (Zarebska et al, 2013: 2901).
Elite power athletes were more likely to possess the CC genotype; 40 percent of power athletes had the genotype whereas 13 percent of endurance had it and 18 percent of non-athletes had it. So power athletes were more than three times as likely to have the CC genotype, compared to endurance athletes and twice as likely to have it compared to non-athletes. On the other hand, one copy of the C allele was found in 55 percent of the power athletes whereas, for the endurance athletes and non-athletes, the C allele was found in about 40 percent of individuals. (Further, in the elite anaerobic athlete, explosive power was consistently found to be a difference maker in predicting elite sporting performance; Lorenz et al, 2013.)
Now we come to the more interesting parts: ethnic differences in the M235T polymorphism. Zarebska et al (2013: 2901-2902) write:
The M235T allele distribution varies widely according to the subject’s ethnic origin: the T235 allele is by far the most frequent in Africans (;0.90) and in African-Americans (;0.80). It is also high in the Japanese population (0.65–0.75). The T235 (C4027) allele distribution of the control participants in our study was lower (0.40) but was similar to that reported among Spanish Caucasians (0.41), as were the sports specialties of both the power athletes (throwers, sprinters, and jumpers) and endurance athletes (marathon runners, 3- to 10-km runners, and road cyclists), thus mirroring the aforementioned studies.
Zarebska et al (2013: 2902) conclude that their study—along with the study they replicated—supports the hypothesis that the C allele of the M235T polymorphism in the AGT gene may confer a competitive advantage in power-oriented sports, which is partly mediated through ANGII production in the skeletal muscles. Mechanisms can explain the mediation of ANGII production in skeletal muscles, such as a direct skeletal muscle hypertrophic effect, along with the redistribution of between muscle blood flow between type I (slow twitch) and II fibers (fast twitch), which would then augment power and speed. However, it is interesting to note that Zarebska et al (2013) did not find any differences between “top-elite” level athletes who had won medals in international competitions compared to elite-level athletes who were not medalists.
The big deal about this gene is that the AGT gene is part of the renin-angiotensin system which is partly responsible for blood pressure and body salt regulation (Hall, 1991; Schweda, 2014). There seems to be an ethnic difference in this polymorphism, and, according to Zarebska et al (2013), African Americans and Africans are more likely to have the polymorphisms that are associated with elite power performance.
There is also a meta-analysis on genotyping and elite power athlete performance (Weyerstrab et al, 2017). Weyerstrab et al (2017) meta-analyzed 36 studies which attempted to find associations between genotype and athletic ability. One of the polymorphisms studied was the famous ACTN3. It has been noted that, when conditions are right (i.e., the right morphology), the combined effects of morphology along with the contractile properties of the individual muscle fibers contribute to the enhanced performance of those with the RR ACTN3 genotype (Broos et al, 2016), while Ma et al (2013) also lend credence to the idea that genetics influences sporting performance. This is, in fact, the most-replicated association in regard to elite sporting performance: we know the mechanism behind how muscle fibers contract; we know how the fibers contract and the morphology needed to maximize the effectiveness of said fast twitch fibers (type II fibers). (Blacks have a higher proportion of type II fibers [see Caeser and Henry, 2015 for a review].)
Weyerstrab et al (2017) meta-analyzed 35 articles, finding significant associations with genotype and elite power performance. They found that ten polymorphisms were significantly associated with power athlete states. Their most interesting findings, though, were on race. Weyerstrab et al (2017: 6) write:
Results of this meta-analysis show that US African American carriers of the ACE AG genotype (rs4363) were more than two times more likely to become a power athlete compared to carriers of the ACE preferential genotype for power athlete status (AA) in this population.
“Power athlete” does not necessarily have to mean “strength athlete” as in powerlifters or weightlifters (more on weightlifters below).
Lastly, the AGT M235T polymorphism, while associated with other power movements, was not associated with elite weightlifting performance (Ben-Zaken et al, 2018). As noted above, this polymorphism was observed in other power athletes, and since these movements are largely similar (short, explosive movements), one would rightly reason that this association should hold for weightlifters, too. However, this is not what we find.
Weightlifting, compared to other explosive, power sports, is different. The beginning of the lifts take explosive power, but during the ascent of the lift, the lifter moves the weight slower, which is due to biomechanics and a heavy load. Ben-Zaken et al (2018) studied 47 weightlifters (38 male, 9 female) and 86 controls. Every athlete that was studied competed in national and international meets on a regular basis. Thirty of the weightlifters were also classified as “elite”, which entails participating in and winning national and international competitions such as the Olympics and the European and World Championships).
Ben-Zaken et al (2018) did find that weightlifters had a higher prevalence of the AGT 235T polymorphism when compared to controls, though there was no difference in the prevalence of this polymorphism when elite and national-level competitors were compared, which “[suggests] that this polymorphism cannot determine or predict elite competitive weightlifting performance” (Ben-Zaken et al, 2018: 38). Of course, a favorable genetic profile is important for sporting success, though, despite the higher prevalence of AGT in weightlifters compared to controls, this could not explain the difference between national and elite-level competitors. Other polymorphisms could, of course, contribute to weightlifting success, variables “such as training experience, superior equipment and facilities, adequate nutrition, greater familial support, and motivational factors, are crucial for top-level sports development as well” (Ben-Zaken et al, 2018: 39).
I should also comment on Anatoly Karlin’s new article The (Physical) Strength of Nations. I don’t disagree with his main overall point; I only disagree that grip strength is a good measure of overall strength—even though it does follow the expected patterns. Racial differences in grip strength exist, as I have covered in the past. Furthermore, there are associations between muscle strength and longevity, with stronger men being more likely to live longer, fuller lives (Ruiz et al, 2008; Volkalis, Haille, and Meisinger, 2015; Garcia-Hermosa, et al, 2018) so, of course, strength training can only be seen as a net positive, especially in regard to living a longer and fuller life. Hand grip strength does have a high correlation with overall strength (Wind et al, 2010; Trosclair et al, 2011). While handgrip strength can tell you a whole lot about your overall health (Lee et al, 2016), of course, there is no better proxy than actually doing the lifts/exercises to ascertain one’s level of strength.
There are replicated genetic associations between explosive, powerful athletic performance, along with even the understanding of the causal mechanisms behind the polymorphisms and their carry-over to power sports. We know that if morphology is right and the individual has the RR ACTN3 genotype, that they will exceed in explosive sports. We know the causal pathways of ACTN3 and how it leads to differences in sprinting competitions. It should be worth noting that, while we do know a lot more about the genomics of sports than we did 20, even 10 years ago, current genetic testing has zero predictive power in regard to talent identification (Pitsladis et al, 2013).
So, of course, for parents and coaches who wonder about the athletic potential of their children and students, the best way to gauge whether or not they will excel in athletics is…to have them compete and compare them to other kids. Even if the genetics aspect of elite power performance is fully unlocked one day (which I doubt it will be), the best way to ascertain whether or not one will excel in a sport is to put them to the test and see what happens. We are in our infancy in understanding the genomics of sporting performance, but when we do understand which genotypes are more prevalent in regard to certain sports (and of course the interactions of the genotype with the environment and genes), then we can better understand how and why others are better in certain sports.
The genomics of elite sporting performance is very interesting; however, the answer that reductionists want to see will not appear: genes are difference makers (Sterelny and Griffith, 1999), not causes, and along with a whole slew of other environmental and mental factors (Lippi, Favaloro, and Guidi 2008), along with a favorable genetic profile with sufficient training (and everything else that comes along with it) are needed for the athlete to reach their maximum athletic potential (see Guth and Roth, 2013). Genetic and environmental differences between individuals and groups most definitely explain differences in elite sporting performance, though elucidating what causes what and the mechanisms that cause the studied trait in question will be tough.
Just because group A has gene or gene networks G and they compete in competition C does not mean that gene or gene networks G contribute in full—or in part—to sporting success. The correlations could be coincidental and non-functional in regard to the sport in question. Athletes should be studied in isolation, meaning just studying a specific athlete in a specific discipline to ascertain how, what, and why works for the specific athlete along with taking anthropomorphic measures, seeing how bad they want “it”, and other environmental factors such as nutrition and training. Looking at the body as a system will take us away from privileging one part over another—while we also do understand that they do play a role but not the role that reductionists believe.
These studies, while they attempt to show us how genetic factors cause differences at the elite level in power sports, they will not tell the whole story, because we must look at the whole system, not reduce it down to the sum of its parts (Shenk, 2011: chapter 5). While blacks are more likely to have these polymorphisms that are associated with elite power athlete performance, this does not obviously carry over to strongman and powerlifting competition.
Height and IQ Genes
1100 words
Genes account for about 80 percent of the variation in height and IQ, with both height and IQ correlating at .2. Therefore, genes must contribute largely to population variances in height. However, finding certain genes that contribute largely to these two traits is a problem, largely because both traits are polygenic in nature. Recent research has shown that most—or all–genes are height genes. If this is the case, are most—or all—genes IQ genes?
Height is around 80-90 percent heritable (Peeters et al, 2009). What this means is that the difference between the tallest and shortest 5 percent of the population is 11 inches, with 10 inches being accounted for by genes and 1 inch being accounted for by environment (Heine, 2017: 30). The gene that contributes the most to human height has been found to give 1/6th of an inch (Weedon et al, 2007). However, a recent meta-analysis shows that certain rare alleles give as much as 8/10ths of an inch (Hirschhorn, Deloukas, and Lettre, 2017). Furthermore, thousands of gene variants combined explain about 50 percent of human height (Yang et al, 2010). Yang et al (2010) also found 294,831 SNPs related to people’s height, which is—more or less—12 times the number of genes in our genome (Heine, 2017: 30; the number of genes in our genome is in the range of 19,000-20,000; Ezkurdia et al, 2014). Another meta-analysis found that 697 genetic variants explain about 20 percent of the genetic variation (Wood et al, 2014). Furthermore, according to geneticist David Goldstein, “most genes are height genes” (Goldstein, 2009).
Author of the book DNA is not Destiny and cultural and social psychologist Steven J. Heine writes:
“This means if you wanted to genetically engineer a designer baby who you would like to grow up to be tall, you would have to make almost 300,000 genetic alterations to the genome and you still would only be half way there. When the genetic evidence suggests that almost all genes are related to height, then in a way, we learn close to nothing about the genetic basis of height.” (Heine, 2017: 30)
Hirschhorn, Deloukas, and Lettre, (2017) found 83 rare and low-frequency genes that explain 1.7 percent of the adult heritability of height, along with newly identified and novel variants that explained 2.4 percent, “and all independent variants, known and novel together explained 27.4% of heritability. By comparison, the 697 known height SNPs explain 23.3% of height heritability in the same dataset (vs. 4.1% by the new height variants identified in this ExomeChip study)” (pg 7). So 27.4 percent of the variance is explained by known common variants and these new variants discovered.
Americans who drink more milk are, on average, half an inch taller than Americans who don’t recall drinking as much milk, even after controlling for race, income, and education (Wiley, 2005). This shows the importance milk has on skeletal muscle growth. This increase has even been noticed in Japan, where they increased their milk intake using school lunch programs (Takahasi, 1984), which increased their height by 4 inches (Funatogawa et al, 2009).
We also grow more in the spring and summer than in the fall and winter. This is due to ultraviolet radiation from the sun’s rays that synthesize some of the vitamin D we drink that is in the cow’s milk. Clearly, environmental factors (UV rays, milk consumption, overall nutrition, etc) all have a part to play in human height variation (Heine, 2017: 30). However, if all genes may be height genes, may all genes be IQ genes?
In regards to IQ, 3 genetic variants explain .3 IQ points (Rietvald et al, 2014):
After adjusting the estimated effect sizes of the SNPs (each R2 ∼ 0.0006) for the winner’s curse, we estimate each as R2 ∼ 0.0002 (SI Appendix), or in terms of coefficient magnitude, each additional reference allele for each SNP is associated with an ∼0.02 SD increase in cognitive performance [or 0.3 points on the typical intelligence quotient (IQ) scale].
This is the gene with the highest known effect that we currently know of. No “but undiscovered X means Y!!”, because science isn’t based on ‘what ifs’.
To predict one’s intelligence, you would need all genes on an SNP chip—which contains about 500,000 SNPs—to be able to predict half of the individual variation in IQ (Davies et al, 2011; Chabris et al, 2012; Heine, 2017: 175). Just as is the case with height, it seems that it’s possible that most—if not all—genes are IQ genes.
So, clearly, intelligence is highly polygenic, and, contrary to what Plomin says, it’s doubtful that we’ll be able to genotype one to guesstimate their intelligence level.
This is because you need more than 500,000 SNPs on a gene chip and even still, that would only explain half of the variance. So it’s reasonable to assume—as is the case with height—that all genes are IQ genes.
Chabris et al (2012) write:
One SNP, rs2760118 in SSADH (also known as ALDH5A1), exhibited a nominally significant association with g (t = 2.01, p = .04), but this association did not survive a Bonferroni correction. The mean g values (transformed to the IQ scale) by genotype for this SNP were 98.3, 99.7, and 100.6 for genotypes TT, TC, and CC respectively.
So it seems that all genes are height genes and all genes could possibly be IQ genes (that is, having a small effect). If most genes are height genes, and height is linked to IQ, then most genes should be IQ genes as well. Therefore, it is plausible that all genes are IQ genes.
Finally, I need to talk about the study that everyone is talking about, the study that found 52 new genes for intelligence (Sniekers et al, 2017). However, Razib Khan cautions: “My plain words are this: do not trust, and always verify“. A Google search for “gene found for” brings up 26,300,000 hits. As can be seen with the study that was published the other day on the supposed ‘new hominin’ found in Europe, science journalists use fancy and catchy headlines. “Genes for ___ and ___” is a bad way to put it—few traits are caused by a single gene, and most traits are highly polygenic, height and IQ included.
Do I think we’ll disentangle the intricacies involved with height and IQ? One day. But since at the moment, 500,000 SNPs need to be loaded on a gene chip to explain half of the variation in individual IQ.
Since most—or all—genes are related to height and the same may be so for IQ, we don’t really learn anything knowing the genes that control for these two traits. In regards to Heine’s (2017) example of genetic engineering 300,000 SNPs for height and you’d only be halfway there, I’d assume the same would be true for IQ. Both traits are highly polygenic, with thousands of genes controlling these traits. Genetic engineering a human for high intelligence or height looks to be a long shot—at least until far into the future.
Genetic Reasons for Human Migration
800 words
It’s always talked about in race realist circles what caused the migration Out of Africa and what gives us an urge and sense of wonder for adventure. Well, like with most things in life, there is a genetic reason behind it.
So I use this website called StumbleUpon and I was Stumbling in the Genetics section, and I came across this article, and as I was reading it, I knew that it was going to be rare in Africans.
And what do you know, I was right.
Seems like the gene developed around 40,000 years ago. It’s obviously not just a coincidence or ‘correlation doesn’t equal causation’ argument, as a whole bunch of research backs this gene, and specific alleles, to certain traits in humans, compared to migration, or lack thereof, around the world.
From some research, I found that the abstract of the article says: it’s linked to hyperactivity, novelty-seeking and risk-taking behaviors. Consistent with findings in animal studies was: having more exploratory behavior as well as increased speed and locomotion. Populations with a history of migration in the past 1 to 30,000 years will show the highest instance of having the DRD4 alleles. Those that migrated more miles have a higher rate than those that migrated less. Sedentary populations show a lower instance of the DRD4 allele.
The abstract also says “After compiling existing data on DRD4 allele frequencies of 2,320 individuals from 39 populations and on the migration pattern of these groups, we found that, compared to sedentary populations, migratory populations showed a higher proportion of long alleles for DRD4.”
A correlation of .85 was found with km traveled and rate of DRD4 allele frequency distributions, which support the hypothesis. Nomadic populations had a 10.4 percent higher rate of DRD4 long alleles than sedentary ones.
Here is a blog post talking about the DRD4-7 genes and how they are involved with dopamine levels in the brain and have to do with motivation and behavior.
It cites a researcher saying that the gene occurs in populations that migrated first and furthest out of Africa are more likely to have the gene.
Another blog talks about how Africans with the 4R allele stayed in Africa whereas those with the 7R left and explored the world.
Research has found that it’s easier for the 2R allele to mutate into a 7R allele than for a 7R allele to mutate back to a 4R allele.
So it’s thought that those who emigrated but remained closer to Africa, such as Asia, lost the behavior to explore and settled rather than explore the Americas (which Siberians did do, who eventually became Indians. I would assume that they would have a higher prevalence of this gene, being as they migrated to the Americas while the Africans stayed in Africa, as well as Indians being the furthest genetically from Africans, it also makes sense there). Also saying that those who emigrated to Asia had the 7R, but stayed and it mutated to the 2R over time.
It says looking at the global prevalence of certain allele frequencies, the 4R is more common in Africa, the 2R in Asia, the 7R in the Americas (mostly South), supporting the theory for the 7R allele being the cause for global expansion and positive selection.
Found this article talking about how the DRD4 allele is a role “in the escalation of substance use during adolescence and potential for an enhanced understanding of early-onset substance use.” and those with the 7R allele “Supporting the differential susceptibility to parenting hypothesis, the results suggest a greater preventive effect for youths carrying a 7-repeat allele”.
This NatGeo article says “Most provocatively, several studies tie 7R to human migration. The first large genetic study to do so, led by Chuansheng Chen of the University of California, Irvine in 1999, found 7R more common in present-day migratory cultures than in settled ones. A larger, more statistically rigorous 2011 study supported this, finding that 7R, along with another variant named 2R, tends to be found more frequently than you would expect by chance in populations whose ancestors migrated longer distances after they moved out of Africa. Neither study necessarily means that the 7R form of the gene actually made those ancestors especially restless; you’d have to have been around back then to test that premise with certainty. But both studies support the idea that a nomadic lifestyle selects for the 7R variant.”
All in all, this shows a genetic reason WHY the migration OoA began. It’s very telling that the DDR7 allele first arose when we migrated OoA.