Home » Heritability
Category Archives: Heritability
Why Heritability Estimates are Flawed: A Conceptual Account
1200 words
Introduction
Heritability estimates have been used as a cornerstone and psychology and genetic research. They are designed to quantify the proportion of phenotypic variance in a population that can be attributed to genetic differences among individuals. We’ve known for a while now that heritability isn’t a measure of genetic strength (Moore and Shenk, 2016), but it’s a population-specific estimate of variance. Here I will provide two a priori arguments (one methodological on twins and the EEA and one theoretical based on Noble’s biological relativity argument). The twin critique shows that the twin researcher’s main assumption (equal environments) does not hold while the biological relativity critique shows that h2 is conceptually invalid. This is why there is a missing heritability problem—it never existed in the first place, and the assumptions twin researchers have are false.
The classical twin method
The CTM compares MZ and DZ twins to attempt to quantify the relative contributions of genes and environment in relation to the origin of trait differences between individuals. Perhaps the biggest assumption of the twin researcher is the equal environments assumption (EEA). The EEA assumes that MZ and DZ twins experience equivalent shared environments.
The EEA seems plausible enough: twins reared together should experience compatible environments, regardless of zygosity. But since DZ twins are more genetically similar than DZ twins so they’ll be more phenotypically similar as well. MZ twins are dressed alike, mistaken for one another, or placed in similar social roles compared to DZ twins which leads to more similar environments. So the shared environmental variance for MZ twins exceeds that for DZ twins, violating the EEA.
Clearly this violation throws a wrench into the logic of the CTM. The formula assumes that the greater similarity in MZ twins stems solely from their genetic identity. But if MZ twins experience more similar environments due to their phenotypic similarity (Fosse, Joseph and Richardson, 2015; Joseph et al, 2015), the difference in correlations between MZs and DZs captures genetic variance and excess environmental similarity. Thus, heritability is overestimated (see eg Bingley, Cappellari, and Tatsiramos, 2023) increasing the effect of genes while masking the effects of the environment—in effect, environment is made to look like genes. Thus, h2 fails to isolate genetic influence as intended. (Note that Grayson 1989 explains this as well, but it seems that it’s just ignored.) Here’s the argument:
(1) The classical twin method assumes that its heritability (h2) estimate (Falconer’s formula) isolates the proportion of phenotypic variance due solely to genetic variance.
(2) For the h2 estimate to isolate genetic variance, the shared environmental variance must be equal for MZ and DZ twins.
(3) MZ twins are more genetically similar than DZ twins.
(4) Genetic similarity between individuals leads to greater similarity in their expressed phenotypic traits, and this phenotypic similarity results in greater similarity in their environmental experiences.
(5) Because MZ twins have greater genetic similarity than DZ twins, and genetic similarity leads to phenotypic similarity, which in turn results in environmental similarity, the shared environmental variance is greater for MZ twins compared to DZ twins.
(6) If the shared environmental variance for MZ twins is greater than that for DZ twins, then the EEA is false because it requires that shared environmental variance be equal for both twin types.
(7) If the EEA is false, then we cannot logically infer genetic conclusions from h2, and thus h2 reflects shared environmental variance (c2), rather than genetic variance.
(8) Any method that relies on an assumption that’s logically inconsistent with the principles governing it’s variables – like the relationship between genetic similarity, environmental similarity and phenotypic similarity – cannot accurately isolate its intended causal component and is therefore conceptually untenable.
(9) Thus, the classical twin method is conceptually and logically untenable since it depends on the EEA which, when false, renders h2 a measure of environmental—not genetic—variance.
The biological relativity critique against h2
This argument is theoretical as opposed to methodological, and it relies on Noble’s (2012) biological relativity argument, where there is no privileged level of causation in biological systems. Genes, cells, tissues, organs, organisms, and the environment form an interdependent network where each level influences and is influenced by the other levels. Phenotypes arise from the interaction between all of these levels, not just due to the independent action of any one of the resources.
Heritability rests on a reductionist assumption—that phenotypic variance can be neatly partitioned into genetic and environmental components with genetic effects isolated as a distinct and quantifiable entity. This framework, clearly, privileges the genetic level treating it as separate from the broader biological and ecological context. But Noble’s argument directly contradicts this view. Genes don’t operate in a vacuum and do nothing on their own.
So by attempting to isolate genetic variance, heritability imposed an artificial simplicity on a complex reality (Rose, 2006). Noble’s principle suggests that separation isn’t just an approximation but a fundamental conceptual flaw. Phenotypic variation emerges from the integrated functioning of all biological levels, which then makes it impossible to assign causation to genes alone.
Thus, h2 is conceptually flawed, since it seeks to measure a genetic contribution that cannot be meaningfully disentangled from the holistic system in which it operates. Obviously the conceptual foundation of h2 contradicts the principle of biological relativity. Since h2 attempts to assign a specific portion of trait variance to genes alone, h2 implicitly privileges the genetic level, suggesting that it can be disentangled from the broader biological system. Noble’s argument denies that this is possible while emphasizing holism and rejecting reductionism. Thus, a priori, h2 estimates are fundamentally flawed because they rest on a reductionist framework which assumes a separability of causes which is incompatible with the holistic, relativistic nature of biological causation. Here’s the argument:
(1) Biological relativity holds that there is no privileged level of causation in biological systems: all levels (genes, cells, tissues, organs, organisms, environments) are interdependent in producing phenotypes.
(2) h2 assumes that genetic variance can be isolated and quantified as a distinct contributor to phenotypic variance.
(C) Since biological relativity rejects the isolation of genetic effects, h2 is conceptually invalid as a measure of genetic influence.
Conclusion
Both of these arguments show the same thing—h2 is a deeply flawed concept. The EEA critique exposes a methodological weakness: since MZ twins experience more similar environments than DZ twins, the excess environmental similarity experienced by MZs masquerades as genetic influence, leaving h2 incapable of isolating genetic variance.
But Noble’s biological relativity argument strikes at a deeper conceptual flaw in this practice, since it challenges the theoretical aspects of h2 itself. Since it highlights the interdependence of biological systems, it dismantles the reductionist notion that genetic effects can be separated from other levels of causation. The gene-centric assumption is at ends with the reality of phenotypes being emergent properties of multi-level interactions, which then renders the concept of h2 conceptually incoherent. Therefore, h2 isn’t only empirically questionable but it is theoretically untenable. The conceptual model is just not sound due to how genes really work (Burt and Simon, 2015)
Thus, again, hereditarianism fails conceptually. Even their main “tool” fails for a modicum of reasons not least the main theoretical killshot for heritability estimates—the principle of biological relativity. The reductionist hereditarian paradigm is conceptually and logically untenable, it’s time to throw it away, it’s time to throw it to the dustbin of history.
Strengthening my Argument to Ban IQ Tests
2500 words
Over three years ago I provided an argument with the ultimate conclusion that IQ tests should be banned. The gist of the argument is that if we believed the hereditarian hypothesis is true and we make policy ascription based on the hereditarian hypothesis and the results that were derived from IQ tests, then a policy could be enacted that would harm a group, and if the policy were enacted, then it would do harm to a group. Thus we should ban whatever led to the policy in question, and so if IQ tests led to the policy in question then IQ tests should be banned. In this article, I will strengthen each premise and then I will provide another argument for why IQ tests should be banned. Here’s the argument:
(P1) The Hereditarian Hypothesis is false
(P2) If the Hereditarian Hypothesis is false and we believed it to be true, then policy A could be enacted.
(P3) If Policy A is enacted, then it will do harm to group G.
(C1) If the Hereditarian Hypothesis is false and we believed it to be true and policy A is enacted, then it will do harm to group G (Hypothetical Syllogism, P2, P3).
(P4) If the Hereditarian Hypothesis is false and we believed it to be true and it would harm group G, then we should ban whatever led to policy A.
(P5) If Policy A is derived from IQ tests, then IQ tests must be banned.
(C2) Therefore, we should ban IQ tests (Modus Ponens, P4, P5).
Premise 1: The truth or falsity of this premise would divide people. On the one hand, there are proponents of the hereditarian hypothesis who believe that the hereditarian hypothesis is true, and so by banning their main “measurement tool”, then we would be censoring “the truth of human biodiversity.” But what entails “the hereditarian hypothesis”? The hereditarian hypothesis can also be called the generic theory of intelligence. It’s main claim is that the observed differences in IQ between groups and individuals are largely attributed to genetic factors. For example, Rushton and Jensen (2005) claim to take the middle ground in arguing that it’s 50/50 genes and environment that lead to the IQ phenotype. But Rushton and Jensen (2005: 279) claim that the 50/50 estimate of heritability is too low—80 percent G and 20 percent E is what we should assume:
A conundrum for theorists of all persuasions, however, is that there is too little evidence of any environmental effects. The hereditarian model of Black–White IQ differences proposed in Section 2 (50% genetic and 50% environmental), far from precluding environmental factors, requires they be found. Although evidence in Sections 3 to 11 provided strong support for the genetic component of the model, evidence from Section 12 was unable to identify the environmental component. On the basis of the present evidence, perhaps the genetic component must be given greater weight and the environmental component correspondingly reduced. In fact, Jensen’s (1998b, p. 443) latest statement of the hereditarian model, termed the default hypothesis, is that genetic and cultural factors carry the exact same weight in causing the mean Black–White difference in IQ as they do in causing individual differences in IQ, about 80% genetic–20% environmental by adulthood.
I have spent the better part of 3 years since publishing my original article to ban IQ tests arguing against the falsity of the hereditarian hypothesis on many grounds. The hereditarian hypothesis largely relies on heritability estimates derived from twin and adoption studies (and now shifting to neuroscience, like they have been since the 80s) and this is where the “laws of behavioral genetics” came from, but the “laws” fail. Important for the hereditarian position is the claim that science can study the mind. However, science is third-personal while mind is first-personal and subjective. Thus it follows that what is third-personal cannot study what is first-personal. Most important for the hereditarian position is the irreducibility of the mental—for if the claim is that the hereditarian hypothesis is true, then the mental would need to reduce to the physical. Humans have minds which means we have the ability for intentional states and propositional attitudes which implies that humans aren’t fully physical. If the argument there holds then science can’t study what’s immaterial, so there is a part of our constitution that can’t be studied by science. So at the end of the day, the hereditarian hypothesis is a physicalist position on the mind-body problem, but empirical evidence is irrelevant to conceptual arguments so the hereditarian position can’t help us understand the mind-body problem since it is an empirical position based on a supposed relationship between mind (“IQ”) and genes/brain/brain structure. Finally, the claim that there is a “general intelligence” is false; we don’t need a nonexistent, reified thing to explain the intercorrelations on IQ scores between individuals and groups. IQ tests are mere knowledge tests—and since knowledge is class-dependent, then different classes have different psychological and cultural tools, and so they would have different knowledge. Basically, IQ is an arbitrary notion especially due to the fact that tests can and have changed in the past for different social groups like men and women (Rosser, 1989), and two white South African groups (Hilliard, 2012) while Kidder and Rosner (2002) showed unconscious bias in the SAT favoring whites due to how the questions were selected. All of these considerations combine to show that the hereditarian hypothesis is false and that we should not accept conclusions from anyone who uses the hereditarian hypothesis as a guide.
Premise 2: But if we believed the hereditarian hypothesis to be true even when it’s false, then we may harm a group. For example, Jensen espoused some eugenic-type ideas in his infamous 1969 paper, stating:
“Is there a danger that current welfare policies, unaided by eugenic foresight, could lead to the genetic enslavement of a substantial segment of our population?” – Jensen, 1969: 95, How Much Can We Boost IQ and Scholastic Achievement?
“What the evidence on heritability tells us is that we can, in fact, estimate a person’s genetic standing on intelligence from his score on an IQ test.” – Jensen, 1970, Can We and Should We Study Race Difference?
“… the best thing the black community could do would be to limit the birth-rate among the least-able members, which of course is a eugenic proposal.” – A Conversation with Arthur Jensen, American Reinnasance, 1992
What Jensen wrote in his 1969 paper is similar to what Herrnstein and Murray (1994: 548) wrote:
We can imagine no recommendation for using the govemment to manipulate fertility that does not have dangers. But this highlights the problem: The United States already has policies that inadvertently social-engineer who has babies, and it is encouraging the wrong women. If the United States did as much to encourage high-IQ women to have babies as it now does to encourage low-lQ women, it would rightly be described as engaging in aggressive manipulation of fertility. The technically precise description of America’s fertility policy is that it subsidizes births among poor women, who are also disproportionately at the low end of the intelligence distribution. We urge generally that these policies, represented by the extensive network of cash and services for low-income women who have babies, be ended.
While these propositions don’t directly stem from hereditarian ideas, they are a direct consequence of such thinking. Like Shockley and Cattell’s beliefs and how their a priori racist ideas influenced the “science” they performed. So premises 2 and 3 presume a causal link between the hereditarian hypothesis, policy A and harm to group G. One specific example that immediately comes to mind is the sterilization or “morons”, “idiots”, and “imbeciles” in the 1900s even continuing up until the late 1970s. Perhaps the most famous case of this was the case of Carrie Buck, to which a judge famously stated, “Three generations of imbeciles are enough.” Premise 3 clearly has historical support.
Conclusion 1: So, since I’ve argued that P2 and P3 are true, then it follows that C1 is true as well. In the original article, I showed that blacks were disproportionately affected by IQ test rulings. Along with the fact that low IQ people were sterilized, this provides yet more support for the premises and the conclusion of this part of the argument.
Premise 4: I have already given the example above about the eugenics movement of the 1900s in America sterilizing thousands of people for having low IQs (this also occurred around the world). The Tuskegee Syphilis Experiment also lends credence to this premise. The US Public Health Service conducted a study from 1932 to 1972 on black Americans where they were observed with syphilis but they weren’t treated after penicillin became available. Segregation laws were based on the belief that the races were inherently different and shouldn’t mix. So in an attempt to prevent mixing, segregation was based on a false belief that blacks were inferior to whites. This is what Darby and Rury (2018) refer to as “the color of mind.”
The Color of Mind [the idea that”blacks were not equal to whites in intelligence, character, or conduct”] has served to rationalize racially exclusionary school practices and unequal educational opportunities, and the effects of these..have worked to sustain this racial ideology
Premise 5: Furthermore, government policies such as redlining and discriniminatory housing policies have led to segregation and inequalities/inequities in education (Rothstein, 2017). These example lend credence to the claim in P4 and P5—policies and practices derived from IQ or other standardized tests can be harmful if they contribute to existing inequalities and disparities. It is quite clear that IQ tests have been used to justify discriminatory polices in the past. Historical and recent considerations point to the fact that IQ tests can and have been used to perpetuate harm on individuals and groups (with the best example being the eugenics movement sterilizing low IQ people, sometimes without their knowledge). The other considerations that weren’t directly related to IQ tests like Tuskegee and Japanese Americans in WW2 show that beliefs that are false that are held to be true can and do lead to devestating consequences for groups of people. The arbitrariness of IQ can also be seen with the death penalty—there are literally life or death consequences riding on the results of a biased test. Moreover, IQ tests have been used to bar immigrants into America in the 1920s (Gould, 1981; Allen, 2006; Richardson, 2011).
Even if IQ tests haven’t been used to enact harmful policies in the past (they quite obviously have), potential future harm is enough. For example, IQ tests are biased in virtue of their item content. So if, say, an employer decides to use IQ tests to select job applicants, they will be necessarily biased by race and class. (Even though IQ tests don’t really have any predictive power for job performance, and whatever relationship between school performance is built in due to the relationship between the items on the tests.)
Thus, the conclusion of the argument that we should ban IQ tests follows. I have argued for the truth of premises 4 and 5 so it then follows that we should ban IQ tests. The argument is valid and I hold it to be sound. So we should ban IQ tests. Nevertheless, here is another argument that we should ban IQ tests:
P1: If IQ tests are not culturally biased and do not perpetuate social inequalities, then they should not be banned.
P2: IQ tests are culturally biased and perpetuate social inequalities.
P3: If IQ tests are culturally biased and perpetuate social inequalities, then IQ tests should be banned.
C: Therefore IQ tests should be banned.
Conclusion
I defended the premises in my original argument more in depth, giving more examples go each premise to justify and strengthen the overall argument. I then gave a new argument stating that since IQ tests at culturally biased, and perpetuate social inequalities then they should be banned. I will now close with a final argument that we should ban IQ tests (hypothetical syllogism):
P1: If IQ tests are biased and have a negative impact on people’s lives, then they should be banned.
P2: If IQ tests are banned, then they will no longer have a negative impact on people’s lives.
C: Therefore, if IQ tests are biased and have a negative impact on people’s lives, then IQ tests should be banned to eliminate that harm.
All you need to do to see the goal of IQ-ists is to merely read what they write. IQ-ists like Jensen and Lynn have outright stated that we should in Jensen’s case limit the birthrate of the “least-able” while there is a danger that “current welfare policies unaided by eugenic foresight” could lead to a “genetic enslavement” of a substantial portion of the population. While in Lynn’s case, he was much more coy about it that we need to “phase out” such cultures (but he claimed it isn’t genocidal, though the term “phase out” of course tells you his real aims). Nevertheless, IQ-ists like Jensen, Lynn, Shockley, and Cattell have told us exactly what their views are. And their views are derived from, ultimately, heritability estimates derived from research with false assumptions. There is also the case that Pygmalion seems to be in the genes—the act of classifying one based on their polygenic score could have feedback effects based on how they view themselves and how society views them: “Through possible mechanisms of stigma and self-fulfilling prophecies, our results highlight the potential psychosocial harms of exposure to low-percentile polygenic scores for educational attainment” (Matthews et al, 2021).
I don’t even think it makes sense to claim that genes contribute to the ontogeny and differences in psychological traits between individuals. Genes only contribute to physical traits. Genes also don’t work how hereditarians need them to work. This is yet another reason why we should reject the hereditarian hypothesis and, along with it, stop using and banning IQ tests. The claim that genes contribute to the differences in psychological traits between people is not only false, but it has caused much harm since the argument has been mounted. Hereditarians have a ton of work to do on the conceptual front if they ever hope to have a sound basis for their beliefs. I’ve argued for a long time that it’s just not possible.
I don’t think we need a moratorium on these matters, such as behavioral genetics. I will be much more specific:
We need to outright cease and ban behavioral genetic research and IQ testing since they lead to avoidable harms. Since these things are based on flawed assumptions, and since these hardly have an evidentiary basis, the only recourse we should take on the matter is to outright ban them. The arguments given here definitively show that to be the case. If someone tells you who they are, then you listen to them. The main actors in the hereditarian sphere have told us who they are and what they stand for for decades, so we should listen to them and ban behavioral genetic and IQ tests. It’s only right to do so.
Hereditarianism is not a Valid Science
1350 words
For years I have been arguing that hereditarianism just isn’t tenable due to the fact that the mental is irreducible to the physical. Since the mental is irreducible to the physical, then hereditarianism cannot possibly be true. I have given many conceptual arguments (here, here, here and here) which argue for (a form of) dualism, and so if dualism is true, then hereditarianism can’t possibly be true.
Here is another argument against hereditarianism:
P1: If hereditarianism is a valid science, then it must be based on a physicalist and reductionist theory of mind.
P2: The mental is irreducible to the physical.
P3: Hereditarianism is based on a physicalist and reductionist theory of mind.
C: Thus, hereditarianism is not a valid science.
Premise 1: The whole hereditarian programme assumes that psychology reduces to genes, which we can see from GWA studies of “intelligence” and other psychological traits. It’s a programme that attempts to show that differences in genes in populations lead to differences in psychological traits. However, this is merely a conceptual confusion.
Since hereditarianism attempts to reduce psychological traits to genes, then it necessarily is a physicalist and reductionist theory of mind. Hereditarianism assumes that actions and behaviors can be reduce to genes, and that we can use the methods they propose to discover these relationships. Hereditarianism, though, is said to be a scientific hypothesis and so it needs testable and falsifiable theories. But, the assumption that psychology reduces to genes is a conceptual one, and so, hereditarianism attempts to make the mind-body problem a scientific problem when it in all actuality is a conceptual argument, to which empirical evidence is irrelevant to.
Hereditarian theorists claim that standardized tests are measurement tools, and so we can then measure and quantify intelligence by administering these tests. However, there is no specified measured object, object of measurement and measurement unit for IQ (Nash, 1990), and for there to be, IQ and whatever other psychological trait the hereditarian claims to be measuring need to have those three things articulated. On another note, hereditarianism would seem to fall prey to a version of what Deacon (1990: 201) calls the numerology fallacy:
Numerology fallacies are apparent correlations that turn out to be artifacts of numerical oversimplification. Numerology fallacies in science, like their mystical counterparts, are likely to be committed when meaning is ascribed to some statistic merely by virtue of its numeric similarity to some other statistic, without supportive evidence from the empirical system that is being described.
Nonetheless, it is clear that when the hereditarian says that the mental can be measured and reduced to genes or brain structure/physiology, they are making a conceptual—not empirical—claim, and so hereditarianism would then fail on conceptual grounds. This is beside the point that (again, conceptually) that there is no a priori privileged level of causation, meaning the gene isn’t a privileged cause over and above other developmental variables (Noble, 2012) and the fact that the conceptual model of heritability and the gene used in hereditarian heritability studies is conceptually flawed (Burt and Simon, 2015). The fact of the matter is, no empirical data can refute these two arguments; these two powerful arguments then combine to refute hereditarianism, making hereditarianism logically untenable.
Hereditarianism must be a physicalist, reductionist account of the mind, and as I have argued for before, this was inevitable. Hereditarianism seeks to either reduce mind to genes or brain structure/physiology, as evidenced by for example Jung and Haier’s (2007) P-FIT model.
Premise 2: I won’t spend much time on this since I have exhaustively argued this claim. But basically, since hereditarianism relies on a physicalist and reductionist account of the mind, then mind either reduces to genes or brain structure/physiology. However, this claim fails conceptually.
Premise 3: This premise states that hereditarianism is a physicalist and reductionist theory of mind. This is evidenced by the fact that since the 80s hereditarians like Richard Haier were attempting to reduce mind (IQ, thinking) to brain physiology using EEG.
Conclusion: It then follows that hereditarianism is not a valid science. No matter how many experiments are carried out by hereditarians, this won’t prove their ideas. The ultimate claim of hereditarianism—and of mind-brain, psychophysical reduction—is a conceptual, not scientific, one.
There is also the fact that the main evidence marshaled for hereditarianism relies on heritability estimates which derive mostly from twin studies. Here’s the argument:
P1: If hereditarianism is a valid science, then it must be based on reliable and valid evidence.
P2: Hereditarianism relies mainly on heritability estimates.
P3: Heritability estimates cannot account for GxE interactions, assume additivity, and can’t account for the complex interactions between G and E.
C: Therefore, hereditarianism cannot be considered a valid science.
Science is based on observation and empirical evidence. Since the advent of twin studies, hereditarianism has relied on heritability estimates, which is a statistical measure of the variance in a trait which can be “explained” by genetic factors. Heritability estimates also assume a heterogeneous environment and that G and E don’t interact. So it then follows that if hereditarianism relies mainly on heritability estimates, then it cannot be a valid science. It doesn’t inform us what the causes of a trait or differences in them are, nor the relative influence of G and E on a trait (Moore and Shenk, 2016). There is also the fact that from these heritability estimates that they have then used and championed GWA studies to find the genes that are causal for differences in IQ scores. However, they would then need to answer the challenge in this article on PGS and I don’t see how anyone can answer it. Nevertheless, “heritability studies attempt the impossible” because “the conceptual biological model on which heritability studies depend—that of identifiably separate effects of genes vs. the environment on phenotype variance—is unsound” (Burt and Simon, 2015).
That hereditarians have shifted to brain imaging and the neurosciences (eg Kirkegaard and Fuerst, 2023) in attempting to validate hereditarianism means I can use the explanatory gap argument to put these newer claims to rest (which is basically the same as the argument I made here against the possibility of science being able to study first-personal subjective states):
P1: Mental states have a first-personal subjective aspect which cannot be captured by third-personal brain sciences.
P2: All physical states can be described in terms of their physical relations relations and properties.
C: So mental states cannot be reduced to third-personal descriptions of brain activity.
So if minds reduce to genes or brains, then we would be able to explain M in terms of P.
P1: If all mental phenomena can be fully explained in terms of physical phenomena, then there is no need for non-physical mental entities or processes.
P2: There are mental phenomena that cannot be fully explained in terms of physical phenomena.
C: Therefore, there are non-physical mental entities or processes.
If physicalism were true, then we would have no need to posit mental entities. But since there are mental phenomena that cannot be fully explained in terms of physical phenomena, then we should accept the existence of non-physical mental phenomena, which would therefore mean that dualism is true and that merely studying brain physiology and processes doesn’t mean that we are studying the mind.
Conclusion
Hereditarianism is hardly a scientific theory. It’s not a scientific theory since M doesn’t reduce to P. It’s not a scientific theory since science can’t study first-personal subjective states. The hereditarian hypothesis cannot be tested in a meaningful way—so it is therefore ad hoc. Hereditarianism should be laid to rest with other hypotheses like phlogiston. Hereditarianism makes no testable predictions. The hereditarian hypothesis is a scientific theory if and only if mind reduces to brain. But the mind doesn’t reduce to the brain. So, again, the hereditarian hypothesis isn’t a scientific theory and, therefore, the mind cannot be studied by science.
Hereditarianism should take it’s place in the annals of failed hypotheses. Hereditarians should stop claiming that hereditarianism is a scientific theory/hypothesis because it very clearly is not.
The Myth of “General Intelligence”
5000 words
Introduction
“General Intelligence” or g is championed as the hallmark “discovery” of psychology. First “discovered” by Charles Spearman in 1904, noting that schoolchildren who scored highly on one test scored highly on others and vice versa for lower-scoring children, he assumed that due to the correlation between tests, that there must be an underlying physiological basis to the correlation, which he posited to be some kind of “mental energy”, stating that the central nervous system (CNS) explained the correlation. He proclaimed that g really existed and that he had verified Galton’s claim of a unitary general ability (Richardson, 2017: 82-83). Psychometricians then claim, from these intercorrelations of scores, that what is hidden from us is then revealed, and that the correlations show that something exists and is driving the correlation in question. That’s the goal of psychometrics/psychology—to quantify and then measure psychological traits/mental abilities. However, I have argued at length that it is a conceptual impossibility—the goal of psychometrics is an impossibility since psychometrics isn’t measurement. Therefore, claims that IQ tests measure g is false.
First, I will discuss the reification of g and it’s relation to brain properties. I will argue that if g is a thing then it must have a biological basis, that is it must be a brain property. Reductionists like Jensen have said as much. But it’s due to the reification of g as a concrete, physical thing that has people hold such beliefs. Second, I will discuss Geary’s theory that g is identical with mitochondrial functioning. I will describe what mitochondria does, and what powers it, and then discuss the theory. I will have a negative view of it, due to the fact that he is attempting to co-opt real, actual functions of a bodily process and attempt to weave g theory into it. Third, I will discuss whether or not psychological traits are indeed quantifiable and measurable, and whether or not there is a definition psychometricians can use to ground their empirical investigations. I will argue negatively for all three. Fourth, I will discuss Herrnstein and Murray’s 6 claims in The Bell Curve about IQ and provide a response to each in turn. Fifth, I will discuss the real cause of score variation, which isn’t reduction to a so-called assumed existence of a biological process/mechanism, but which is due to affective factors and exposure to the specific type of knowledge items on the test. Lastly, I will conclude and give an argument for why g isn’t a thing and is therefore immeasurable.
On reifications and brain properties
Contrary to protestations from psychometricians, they in fact do reifiy correlations and then claim that there exists some unitary, general factor that pervades all mental tests. If reification is treating the abstract as something physical, and if psychometrics treat g as something physical, then they are reifying g based on mere intercorrelations between tests. I am aware that, try as they might, they do attempt to show that there is an underlying biology to g, but these claims are defeated by the myriad arguments I’ve raised against the reducibility of the mental to the physical. Another thing that Gould gets at is that psychometricians claim that they can rank people—this is where the psychometric assumption that because we can put a number to their reified thing, that there is something being measured.
Reification is “the propensity to convert an abstract concept (like intelligence) into a hard entity (like an amount of quantifiable brain stuff)” (Gould, 1996: 27). So g theorists treat g as a concrete, physical, thing, which then guides their empirical investigations. They basically posit that the mental has a material basis, and they claim that they can, by using correlations between different test batteries, we can elucidate the causal biological mechanisms/brain properties responsible for the correlation.
Spearman’s theory—and IQ—is a faculty theory (Nash, 1990). It is a theory in which it is claimed that the mind is separated into different faculties, where mental entities cause the intellectual performance. Such a theory needs to keep up the claim that a cognitive faculty is causally efficacious for information processing. But the claim that the mind is “separated” into different faculties fails, and it fails since the mind is a single sphere of consciousness, it is not a complicated arrangement of mental parts. Physicalists like Jensen and Spearman don’t even have a sound philosophical basis on which to ground their theories. Their psychology is inherently materialist/physicalist, but materialism/physicalism is false and so it follows that their claims do not hold any water. The fact of the matter is, Spearman saw what he wanted to see in his data (Schlinger, 2003).
I have already proven that since dualism is true, then the mental is irreducible to the physical and since psychometrics isn’t measurement, then what psychometricians claim to do just isn’t possible. I have further argued that science can’t study first-personal subjective states since science is third-personal and objective. The fact is the matter is, hereditarian psychologists are physicalist, but it is impossible for a purely physical thing to be able to think. Claims from psychometricians about their “mental tests” basically reduce to one singular claim: that g is a brain property. I have been saying this for years—if g exists, it has to be a brain property. But for it to be a brain property, one needs to provide defeaters for my arguments against the irreducibility of the mental and they also need to argue against the arguments that psychometrics isn’t measurement and that psychology isn’t quantifiable. They can assume all they want that it is quantifiable and that since they are giving tests, questionnaires, likert scales, and other kinds of “assessments” to people that they are really measuring something; but, ultimately, if they are actually measuring something, then that thing has to be physical.
Jensen (1999) made a suite of claims trying to argue for a physical basis for g,—to reduce g to biology—though, upon conceptual examination (which I have provided above) these claims outright fail:
g…[is] a biological [property], a property of the brain
The ultimate arbiter among various “theories of intelligence” must be the physical properties of the brain itself. The current frontier of g research is the investigation of the anatomical and physiological features of the brain that cause g.
…psychometric g has many physical correlates…[and it] is a biological phenomenon.
As can be seen, Jensen is quite obviously claiming that g is a biological brain property—and this is what I’ve been saying to IQ-ists for years: If g exists, then it MUST be a property of the brain. That is, it MUST have a physical basis. But for g proponents to show this is in fact reality, they need to attempt to discredit the arguments for dualism, that is, they need to show that the mental is reducible to the physical. Jensen is quite obviously saying that a form of mind-brain identity is true, and so my claim that it was inevitable for hereditarianism to become a form of mind-brain identity theory is quite obviously true. The fact of the matter is, Jensen’s beliefs are reliant upon an outmoded concept of the gene, and indeed even a biologically implausible heritability (Richardson, 1999; Burt and Simons, 2014, 2015).
But Jensen (1969) contradicted himself when it comes to g. On page 9, he writes that “We should not reify g as an entity, of course, since it is only a hypothetical construct intended to explain covariation am ong tests. It is a hypothetical source of variance (individual differences) in test scores.” But then 10 pages later on pages 19-20 he completely contradicts himself, writing that g is “a biological reality and not just a figment of social conventions.” That’s quite the contradiction: “Don’t reifiy X, but X is real.” Jensen then spent the rest of his career trying to reduce g to biology/the brain (brain properties), as we see above.
But we are now in the year 2023, and so of course there are new theoretical developments which attempt to show that Spearman’s hypothesized mental energy really does exist, and that it is the cause of variations in scores and of the positive manifold. This is now where we will turn.
g and mitochondrial functioning
In a series of papers, David Geary (2018, 2019, 2020, 2021) tries to argue that mitochondriaal functioning is the core component in g. At last, Spearman’s hypothetical construct has been found in the biology of our cells—or has it?
One of the main functions of mitochondria is to oxidative phosphorylation to produce adenosine triphosphate (ATP). All living cells use ATP as fuel, it acts as a signaling molecule, it is also involved in cellular differentiation and cell death (Khakh and Burnstock, 2009). The role of mitochondrial functioning in spurring disease states has been known for a while, such as with cardiovascular diseases such as cardiomyopathy (Murphy et al, 2016, Ramaccini at al, 2021).
So due to the positive manifold, where performance in one thing is correlated with a performance in another, Geary assumes—as Spearman and Jensen did before him—that there must be some underlying biological mechanism which then explains the correlation. Geary then uses established outcomes of irregular mitochondrial functioning to then argue that the mental energy that Spearman was looking for could be found in mitochondrial functioning. Basically, this mental energy is ATP. I don’t deny that mitochondriaal functioning plays a role in the acquisition of disease states, indeed this has been well known (eg, Gonzales et al, 2022). What I deny is Gary’s claim that mitochondrial functioning has identity with Spearman’s g.
His theory is, like all other hereditarian-type theories, merely correlative—just like g theory. He hasn’t shown any direct, causal, evidence of mitochondrial functioning in “intelligence” differences (nor for a given “chronological age). That as people age their bodies change which then has an effect on their functioning doesn’t mean that the powerhouse of the cell—ATP—is causing said individual differences and the intercorrelations between tests (Sternberg, 2020). Indeed, environmental pollutants affect mitochondrial functioning (Byun and Baccarelli, 2014; Lambertini and Byun, 2016). Indeed, most—if not all—of Geary’s hypotheses do not pass empirical investigation (Schubert and Hagemann, 2020). So while Geary’s theory is interesting and certainly novel, it fails in explaining what he set out to.
Quantifiable, measurable, definable, g?
The way that g is conceptualized is that there is a quantity of it—where one has “more of it” than other people, and this, then, explains how “intelligent” they are in comparison to others—so implicit in so-called psychometric theory is that whatever it is their tests are tests of, something is being quantified. But what does it mean to quantify something? Basically, what is quantification? Simply, it’s the act of giving a numerical value to a thing that is measured. Now we have come to an impasse—if it isn’t possible to measure what is immaterial, how can we quantify it? That’s the thing, we can’t. The g approach is inherently a biologically reductionist one. Biological reductionism is false. So the g approach is false.
Both Gottfredson (1998) and Plomin (1999) make similar claims to Jensen, where they talks about the “biology of g” and the “genetics of g“. Plomin (1999) claims that studies of twins show that g has a substantial heritability, while Gottfredson (1998) claims that heritability of IQ increases to up until adulthood where it “rises to 60 percent in adolescence and to 80 percent by late adulthood“, citing Bouchard’s MISTRA (Minnesota Study of Twins Reared Apart). (See Joseph 2022 for critique and for the claim that the heritability of IQ in that study is 0 percent.) They, being IQ-ists, of course assume a genetic component to this mystical g. However, there arguments are based on numerous false assumptions and studies with bad designs (and hidden results), and so they must be rejected.
If X is quantitative, then X is measurable. If X is measurable, then X has a physical basis. Psychological traits don’t have a physical basis. So psychological traits aren’t quantitative and therefore not measurable. Geary’s attempt at arguing for identity between g and mitochondrial functioning is an attempt at a specified measured object for g, though his theory just doesn’t hold. Stating truisms about a biological process and then attempting to liken the process with the construct g just doesn’t work; it’s just a post-hoc rationalization to attempt to liken g with an actual biological process.
Furthermore, if X is quantitative, then there is a specified measured object, object of measurement and measurement unit for X. But this is where things get rocky for g theorists and psychometricians. Psychometry is merely pseudo-measurement. Psychometricians cannot give a specified measured object, and if they can’t give a specified measured object they cannot give an object of measurement. They thusly also cannot construct a measurement unit. Therfore, “the necessary conditions for metrication do not exist” (Nash, 1990: 141). Even Haier (2014, 2018) admits that IQ test scores don’t have a unit that is like inches, liters, or grams. This is because those are ratio scales and IQ is ordinal. That is, there is no “0-point” for IQ, like there is for other actual, real measures like temperature. That’s the thing—if you have a thing to be measured, then you have a physical object and consequently a measument unit. But this is just not possible for psychometry. I then wonder why Haier doesn’t follow what he wrote to its logical conclusion—that the project of psychometrics is just not possible. Of course the concept of intelligence doesn’t have a referent, that is, it doesn’t name a property like height, weight, or temperature (Midgley, 2018:100-101). Even the most-cited definition of intelligence—Gottfredson’s—still fails, since she contradicts herself in her very definition.
Of course IQ “ranks” people by their performance—some people perform better on the test than others (which is an outcome of prior experience). So g theorists and IQ-ists assume that the IQ test is measuring some property that varies between groups which then leads to score differences on their psychometric tests. But as Roy Nash (1990: 134) wrote:
It is impossible to provide a satisfactory, that is non-circular, definition of the supposed ‘general cognitive ability’ IQ tests attempt to measure and without that definition IQ theory fails to meet the minimal conditions of measurement.
But Boeck and Kovas (2020) try to sidestep this issue with an extraordinary claim, “Perhaps we do not need a definition of intelligence to investigate intelligence.” How can we investigate something sans a definition of the object of investigation? How can we claim that a thing is measured if we have no definition, and no specified measured object, object of measurement and measurement unit, as IQ-ists seem to agree with? Again, IQ-ists don’t take these conclusions to their further logical conclusion—that we simply just cannot measure and quantify psychological traits.
Haier claims that PGS and “DNA profiles” may lead to “new definitions of intelligence” (however ridiculous a claim). He also, in 2009, had a negative outlook on identifying a “neuro g” since “g-scores derived from different test batteries do not necessarily have equivalent neuro-anatomical substrates, suggesting that identifying a “neuro-g” will be difficult” (Haier, 2009). But one more important reason exists, and it won’t just make it “difficult” to identify a neuro g, it makes it conceptually impossible. That is the fact that cognitive localizations are not possible, and that we reify a kind of average in brain activations when we look at brain scans using fMRI. The fact of the matter is, neuroreduction just isn’t possible, empirically (Uttal, 2001, 2014, 2012), nor is it possible conceptually.
Herrnstein and Murray’s 6 claims
Herrnstein and Murray (1994) make six claims about IQ (and also g):
(1) There is such a thing as a general factor of cognitive on which human beings differ.
Of course implicit in this claim is that it’s a brain property, and that people have this in different quantities. However, the discussion above puts this claim to bed since psychological traits aren’t quantitative. This, of course comes from the intercorrelations of test scores. But we will see that most of the source of variation isn’t even entirely cognitive and is largely affective and due to one’s life experiences (due to the nature of the item content).
(2) All standardized tests of academic aptitude or achievement measure this general factor to some degree, but IQ tests expressly designed for that purpose measure it most accurately.
Of course Herrnstein and Murray are married to the idea that these tests are measures of something, that since they give different numbers due to one’s performance, there must be an underlying biology behind the differences. But of course, psychometry isn’t true measurement.
(3) IQ scores match, to a first degree, whatever it is that people mean when they use the word intelligent or smart in ordinary language.
That’s because the tests are constructed to agree with prior assumptions on who is or is not “intelligent.” As Terman constructed his Stanford-Binet to agree with his own preconceived notions of who is or is not “intelligent”: “By developing an exclusion-inclusion criteria that favored the aforementioned groups, test developers created a norm “intelligent” (Gersh, 1987, p.166) population “to differentiate subjects of known superiority from subjects of known inferiority” (Terman, 1922, p. 656)” (Bazemoore-James, Shinaprayoon, and Martin 2017). Of course, since newer tests are “validated”(that is, correlated with) older, tests (Richardson, 1991, 2000, 2002, 2017; Howe, 1997), this assumption is still alive today.
(4) IQ scores are stable, although not perfectly so, over much of a person’s life.
IQ test scores are malleable, and this of course would be due to the experience one has in their lives which would then have them ready to take a test. Even so, if this claim were true, it wouldn’t speak to the “biology” of g.
(5) Properly administered IQ tests are not demonstrably biased against social, economic, ethnic, or racial groups.
This claim is outright false and can be known quite simply: the items on IQ tests derive from specific classes, mainly the white middle-class. Since this is true, it would then follow that people who are not exposed to the item content and test structures wouldn’t be as prepared as those who are. Thus, IQ tests are biased against different groups, and if they are biased against different groups it also follows that they are biased for certain groups, mainly white Americans. (See here for considerations on Asians.)
(6) Cognitive ability is substantially heritable, apparently no less than 40 percent and no more than 80 percent.
It’s nonsense to claim that one can apportion heritability into genetic and environmental causes, due to the interaction between the two. IQ-ists may claim that twin, family, and adoption studies show that IQ is X amount heritable so there must thusly be a genetic component to differences in test scores. But the issue with heritability has been noted for decades (see Charney, 2012, 2016, 2022; Joseph, 2014, Moore and Shenk, 2016, Richardson, 2017) so this claim also fails. There is also the fact that behavioral genetics doesn’t have any “laws.” It’s simply fallacious to believe that nature and nurture, genes and environment, contribute additively to the phenotype, and that their relative contributions to the phenotype can be apportioned. But hereditarians need to keep that facade up, since it’s the only way their ideas can have a chance at working.
What explains the intercorrelations?
We still need an explanation of the intercorrelations between test scores. I have exhaustively argued that the usual explanations from hereditarianism outright fail—g isn’t a biological reality and IQ tests aren’t a measure at all because psychometrics isn’t measurement. So what explains the intercorrelations? We know that IQ tests are comprised of different items, whether knowledge items or more “abstract” items like the Raven. Therefore, we need to look to the fact that people aren’t exposed to certain things, and so if one comes across something novel that they’ve never been exposed to, they thusly won’t know how to answer it and their score will then be affected due to their ignorance of the relationship between the question and answer on the test. But there are other things irrespective of the relationship between one’s social class and the knowledge they’re exposed to, but social class would still then have an effect on the outcome.
IQ is, merely, numerical surrogates for class affiliation (Richardson, 1999; 2002; 2022). The fact of the matter is, all human cognizing takes place in specific cultural contexts in which cultural and psychological tools are used. This means, quite simply, that culture-fair tests are impossible and, therefore, that such tests are necessarily biased against certain groups, and so they are biased for certain groups. Lev Vygotsky’s sociocultural theory of cognitive development and his concepts of psychological and cultural tools is apt here. This is wonderfully noted by Richardson (2002: 288):
IQ tests, the items of which are designed by members of a rather narrow social class, will tend to test for the acquisition of a rather particular set of cultural tools: in effect, to test, or screen, for individuals’ psychological proximity to that set per se, regardless of intellectual complexity or superiority as such.
Thinking is culturally embedded and contextually-specific (although irreducible to physical things), mediated by specific cultural tools (Richardson, 2002). This is because one is immersed in culture immediately from birth. But what is a cultural tool? Cultural tools include language (Weitzman, 2013) (it’s also a psychological tool), along with “different kinds of numbering and counting, writing schemes, mnemonic technical aids, algebraic symbol systems, art works, diagrams, maps, drawings, and all sorts of signs (John-Steiner & Mahn, 1996; Stetsenko, 1999)” (Robbins, 2005). Children are born into cultural environments, and also linguistically-mediated environments (Vasileva and Balyasnikova, 2019). But what are psychological tools? One psychological tool (which would also of course be cultural tools) would be words and symbols (Vallotton and Ayoub, 2012).
Vygotsky wrote: “In human behavior, we can observe a number of artificial means aimed at mastering one’s own psychological processes. These means can be conditionally called psychological tools or instruments… Psychological tools are artificial and intrinsically social, rather than natural and individual. They are aimed at controlling human behavior, no matter someone else’s or one’s own, just as technologies are aimed at controlling nature” (Vygotsky, 1982, vol. 1, p. 103, my translation). (Falikman, 2021).
The source of variation in IQ tests, after having argued that social class is a compound of the cultural tools one is exposed to. Furthermore, it has been shown that the language and numerical skills used on IQ tests are class-dependent (Brito, 2017). Thus, the compounded cultural tools of different classes and racial groups then coalesce to explain how and why they score the way they do. Richardson (2002: 287-288) writes
that the basic source of variation in IQ test scores is not entirely (or even mainly) cognitive, and what is cognitive is not general or unitary. It arises from a nexus of sociocognitive-affective factors determining individuals’ relative preparedness for the demands of the IQ test. These factors include (a) the extent to which people of different social classes and cultures have acquired a specific form of intelligence (or forms of knowledge and reasoning); (b) related variation in ‘academic orientation’ and ‘self-efficacy beliefs’; and (c) related variation in test anxiety, self-confidence, and so on, which affect performance in testing situations irrespective of actual ability.
Basically, what explains the intercorrelations of test scores—so-called g—are affective, non-cognitive factors (Richardson and Norgate, 2015). Being prepared for the tests, being exposed to the items on the tests (from which are drawn from the white middle-class) explains IQ score differences, not a mystical g that some have more of than others. That is, what explains IQ score variation is one’s “distance” from the middle-class—this follows due to the item content on the test. At the end of the day, IQ tests don’t measure the ability for complex cognition. (Richardson and Norgate, 2014). So one can see that differing acquisition of cultural tools by different cultures and classes would then explain how and why individuals of those groups then attain different knowledge. This, then, would license the claim that one’s IQ score is a mere outcome of their proximity to the certain cultural tools in use in the tests in question (Richardson, 2012).
The fact of the matter is, children do not enter school with the same degree of readiness (Richardson, 2022), and this is due to their social class and the types of things they are exposed to in virtue of their class membership (Richardson and Jones, 2019). Therefore, the explanation for these differences in scores need not be some kind of energy that people have in different quantities, it’s only the fact that from birth we are exposed to different cultures and therefore different cultural and psychological tools which then causes differences in the readiness of children for school. We don’t need to posit any supposed biological mechanism for that, when the answer is clear as day.
Conclusion
As can be seen from this discussion, it is clear that IQ-ist claims of g as a biological brain property fail. They fail because psychometrics isn’t measurement. They fail because psychometricians assume that what they are “measuring” (supposedly psychological traits) have a physical basis and have the necessary components for metrication. They fail because the proposed biology to back up g theory don’t work, and claiming identity between g and a biological process doesn’t mean that g has identity between that biological process. Merely describing facts about physiology and then attempting to liken it to g doesn’t work.
Psychologists try so very hard for psychology to be a respected science, even when what they are studying bares absolutely no relationship to the objects of scientific study. Their constructs are claimed to be natural kinds, but they are merely historically contingent. Due to the way these tests are constructed, is it any wonder why such score differences arise?
The so-called g factor is also an outcome of the way tests are constructed:
Subtests within a battery of intelligence tests are included n the basis of them showing a substantial correlation with the test as a whole, and tests which do not show such correlations are excluded. (Tyson, Jones, and Elcock, 2011: 67)
This is why there is a correlation between all subtests that comprise a test. Because it is an artificial creation of the test constructors, just like their normal curve. Of course if you pick and choose what you want in your battery or test, you can then coax it to get the results you want and then proclaim that what explains the correlations are some sort of unobserved, hidden variable that individuals have different quantities of. But the assumption that there is a quantity of course assumes that there is a physical basis to that thing. Physicalists like Jensen, Spearman, and then Haier of course presume that intelligence has a physical basis and is either driven by genes or can be reduced to neurophysiology. These claims don’t pass empirical and conceptual analysis. For these reasons and more, we should reject claims from hereditarian psychologists when they claim that they have discovered a genetic or neurophysiological underpinning to “intelligence.”
At the end of the day, the goal of psychometrics is clearly impossible. Try as they might, psychometricians will always fail. Their “science” will never be on the level of physics or chemistry, and that’s because they have no definition of intelligence, nor a specified measured object, object of measurement and measurement unit. They know this, and they attempt to construct arguments to argue their way out of the logical conclusions of those facts, but it just doesn’t work. “General intelligence” doesn’t exist. It’s a mere creation of psychologists and how they make their tests, so it’s basically just like the bell curve. Intelligence as an essence or quality is a myth; just because we have a noun “intelligence” doesn’t mean that there really exists a thing called “intelligence” (Schlinger, 2003). The fact is the matter is, intelligence is simply not an explanatory concept (Howe, 1997).
IQ-ist ideas have been subject to an all-out conceptual and empirical assault for decades. The model of the gene they use is false, (DNA sequences have no privileged causal role in development), heritability estimates can’t do what they need them to do, how the estimates are derived rest on highly environmentally-confounded studies, the so-called “laws” of behavioral genetics are anything but, they lack definitions and specified measured objects, objects of measurement and measurement units. It is quite simply clear that hereditarian ideas are not only empirically false, but they are conceptually false too. They don’t even have their concepts in order nor have they articulated exactly WHAT it is they are doing, and it clearly shows. The reification of what they claim to be measuring is paramount to that claim.
This is yet another arrow in the quiver of the anti-hereditarian—their supposed mental energy, their brain property, simply does not, nor can it, exist. And if it doesn’t exist, then they aren’t measuring what they think they’re measuring. If they’re not measuring what they think they’re measuring, then they’re showing relationships between score outcomes and something else, which would be social class membership along with everything else that is related with social class, like exposure to the test items, along with other affective variables.
Now here is the argument (hypothetical syllogism):
P1: If g doesn’t exist, then psychometricians are showing other sources of variation for differences in test scores.
P2: If psychometricians are showing other sources of variation for differences in test scores and we know that the items on the tests are class-dependent, then IQ score differences are mere surrogates for social class.
C: Therefore, if g doesn’t exist, then IQ score differences are mere surrogates for social class.
The “Interactionism Fallacy”
2350 words
A fallacy is an error in reasoning that makes an argument invalid. The “interactionism fallacy” is the fallacy—coined by Gottfredson (2009)—that since genes and environment interact, that heritability estimates are not useful—especially for humans (they are for nonhuman animals where environments can be fully controlled; see Schonemann, 1997; Moore and Shenk, 2016). There are many reasons why this ‘fallacy’ is anything but a fallacy; it is a simple truism: genes and environment (along with other developmental products) interact to ‘construct’ the organism (what Oyama, 2000 terms ‘constructive interactionism—“whereby each combination of genes and environmental influences simultaneously interacts to produce a unique result“). The causal parity thesis (CPT) is the thesis that genes/DNA play an important role in development, but so do other variables, so there is no reason to privilege genes/DNA above other developmental variables (see Noble, 2012 for a similar approach). Genes are not special developmental resources and so, nor are they more important than other developmental resources. So the thesis is that genes and other developmental resources are developmentally ‘on par’.
Genes need the environment. Without the environment, genes would not be expressed. Behavior geneticists claim to be able to partition genes from environment—nature from nurture—on the basis of heritability estimates, mostly gleaned from twin and adoption studies. However, the method is flawed: since genes interact with the environment and other genes, how would it be possible to neatly partition the effects of genes from the effects of the environment? Behavior geneticists claim that we can partition these two variables. Behavior geneticists—and others—cite the “Interactionism fallacy”, the fallacy that since genes interact with the environment that heritability estimates are useless. This “fallacy”, though, confuses the issue.
Behavior geneticists claim to show how genes and the environment affect the ontogeny of traits in humans with twin and adoption studies (though these methods are highly flawed). The purpose of this “fallacy” is to disregard what developmental systems theorists claim about the interaction of nature and nurture—genes and environment.
Gottfredson (2009) coins the “interactionism fallacy”, which is “an irrelevant truth [which is] that an organism’s development requires genes and environment to act in concert” and the “two forces are … constantly interacting” whereas “Development is their mutual product.” Gottfredson also states that “heritability … refers to the percentage of variation in … the phenotype, which has been traced to genetic variation within a particular population.” (She also makes the false claim that “One’s genome is fixed at birth“; though this is false, see epigenetics/methylation studies.) Heritability estimates, according to Phillip Kitcher are “‘irrelevant’ and the fact that behavior geneticists persist
in using them is ‘an unfortunate tic from which they cannot free themselves’ (Kitcher,
2001: 413)” (quoted in Griffiths, 2002).
Gottfredson is engaging in developmental denialism. Developmental denialism “occurs when heritability is treated as a causal mechanism governing the developmental reoccurrence of traits across generations in individuals.” Gottfredson, with her “interactionism fallacy” is denying organismal development by attempting to partition genes from environment. As Rose (2006) notes, “Heritability estimates are attempts to impose a simplistic and reified dichotomy (nature/nurture) on non-dichotomous processes.” The nature vs nurture argument is over and neither has won—contra Plomin’s take—since they interact.
Gottfredson seems confused, since this point was debated by Plomin and Oyama back in the 80s (Plomin’s review of Oyama’s book The Ontogeny of Information; see Oyama, 1987, 1988; Plomin, 1988a, b). In any case, it is true that development requires genes to interact. But Gottfredson is talking about the concept of heritability—the attempt to partition genes and environment through twin, adoption and family studies (which have a whole slew of problems). For example, Moore and Shenk (2016: 6) write:
Heritability statistics do remain useful in some limited circumstances, including selective breeding programs in which developmental environments can be strictly controlled. But in environments that are not controlled, these statistics do not tell us much.
Susan Oyama writes in The Ontogeny of Information (2000, pg 67):
Heritability coefficients, in any case, because they refer not only to variation in genotype but to everything that varied (was passed on) with it, only beg the question of what is passed on in evolution. All too often heritability estimates obtained in one setting are used to infer something about an evolutionary process that occurred under conditions, and with respect to a gene pool, about which little is known. Nor do such estimates tell us anything about development.
Characters are produced by the interaction of nongenetic and genetic factors. The biological flaw, as Moore and Shenk note, throw a wrench into the claims of Gottfredson and other behavior geneticists. Phenotypes are ALWAYS due to genetic and nongenetic factors interacting. So the two flaws of heritability—the environmental and biological flaw (Moore and Shenk, 2016)—come together to “interact” to refute such simplistic claims that genes and environment—nature and nurture—can be separated.
For instance, as Moore (2016) writes, though “twin study methods are among the most powerful tools available to quantitative behavioral geneticists (i.e., the researchers who took up Galton’s goal of disentangling nature and nurture), they are not satisfactory tools for studying phenotype development because they do not actually explore biological processes.” (See also Richardson, 2012.) This is because twin studies ignore biological/developmental processes that lead to phenotypes.
Gamma and Rosenstock (2017) write that the concept of heritability that behavioral geneticists use is “is a generally useless quantity” while “the behavioral genetic dichotomy of genes vs environment is fundamentally misguided.” This brings us back to the CPT; there is causal parity to all processes/interactants that form the organism and its traits, thus the concept of heritability that behavioral geneticists employ is a useless measure. Oyama, Griffiths, and Gray (2001: 3) write:
These often overlooked similarities form part of the evidence for DST’s claim of causal parity between genes and other factors of development. The “parity thesis” (Griffiths and Knight 1998) does not imply that there is no difference between the particulars of the causal roles of genes and factors such as endosymbionts or imprinting events. It does assert that such differences do not justify building theories of development and evolution around a distinction between what genes do and what every other causal factor does.
Behavior geneticists’ endeavor, though, is futile. Aaron Panofsky (2016: 167) writes that “Heritability estimates do not help identify particular genes or ascertain their functions in development or physiology, and thus, by this way of thinking, they yield no causal information.” (Also see Panofsky, 2014; Misbehaving Science: Controversy and the Development of Behavior Genetics.) So, the behavioral genetic method of partitioning genes and environment does not—and can not—show causation for trait ontogeny.
Now, while people like Gottfredson and others may deny it, they are genetic determinists. Genetic determinism, as defined by Griffiths (2002) is “the idea that many significant human characteristics are rendered inevitable by the presence of certain genes.” Using this definition, many behavior geneticists and their sympathizers have argued that certain traits are “inevitable” due to the presence of certain genes. Genetic determinism is literally the idea that genes “determine” aspects of characters and traits, though it has been known for decades that it is false.
Now we can take a look at Brian Boutwell’s article Not Everything Is An Interaction. Boutwell writes:
Albert Einstein was a brilliant man. Whether his famous equation of E=mc2 means much to you or not, I think we can all concur on the intellectual prowess—and stunning hair—of Einstein. But where did his brilliance come from? Environment? Perhaps his parents fed him lots of fish (it’s supposed to be brain food, after all). Genetics? Surely Albert hit some sort of genetic lottery—oh that we should all be so lucky. Or does the answer reside in some combination of the two? How very enlightened: both genes and environment interact and intertwine to yield everything from the genius of Einstein to the comedic talent of Lewis Black. Surely, you cannot tease their impact apart; DNA and experience are hopelessly interlocked. Except, they’re not. Believing that they are is wrong; it’s a misleading mental shortcut that has largely sown confusion in the public about human development, and thus it needs to be retired.
[…]
Most traits are the product of genetic and environmental influence, but the fact that both genes and environment matter does not mean that they interact with one another. Don’t be lured by the appeal of “interactions.” Important as they might be from time to time, and from trait to trait, not everything is an interaction. In fact, many things likely are not.
I don’t even know where to begin here. Boutwell, like Gottfredson, is confused. The only thing that needs to be retired because it “has largely sown confusion in the public about human development” is, ironically, the concept of heritability (Moore and Shenk, 2016)! I have no idea why Boutwell claimed that it’s false that “DNA and experience [environment] are hopelessly interlocked.” This is because, as Schneider (2007) notes, “the very concept of a gene requires an environment.” Since the concept of the gene requires the environment, how can we disentangle them into neat percentages like behavior geneticists claim to do? That’s right: we can’t. Do be lured by the appeal of interactions; all biological and nonbiological stuff constantly interacts with one another.
Boutwell’s claims are nonsense. It would be worth it to quote Richard Lewontin’s forward in the 2000 2nd edition of Susan Oyama’s The Ontogeny of Information (emphasis Lewontin’s):
Nor can we partition variation quantitatively, ascribing some fraction of variation to genetic differences and the remainder to environmental variation. Every organism is the unique consequence of the reading of its DNA in some temporal sequence of environments and subject to random cellular events that arise because of the very small number of molecules in each cell. While we may calculate statistically an average difference between carriers of one genotype and another, such average differences are abstract constructs and must not be reified with separable concrete effects of genes in isolation from the environment in which the genes are read. In the first edition of The Ontogeny of Information Oyama characterized her construal of the causal relation between genes and environment as interactionist. That is, each unique combination of genes and environment produces a unique and a priori unpredictable outcome of development. The usual interactionist view is that there are separable genetic and environmental causes, but the effects of these causes acting in combination are unique to the particular combination. But this claim of ontogenetically independent status of the causes as causes, aside from their interaction in the effects produced, contradicts Oyama’s central analysis of the ontogeny of information. There are no “gene actions” outside environments, and no “environmental actions” can occur in the absence of genes. The very status of environment as a contributing cause to the nature of an organism depends on the existence of a developing organism. Without organisms there may be a physical world, but there are no environments. In like the manner no organisms exist in the abstract without environments, although there may be naked DNA molecules lying in the dust. Organisms are the nexus of external circumstances and DNA molecules that make these physical circumstances into causes of development in the first place. They become causes only at their nexus, and they cannot exist as causes except in their simultaneous action. That is the essence of Oyama’s claim that information comes into existence only in the process of ontogeny. (Oyama, 2000: 16)
There is an “interactionist consensus” (see Oyama, Griffiths, and Grey, 2001; What is Developmental Systems Theory? pg 1-13): the organism and the suite of traits it has is due to the interaction of genetic/environmental/epigenetic etc. resources at every stage of development. Therefore, for organismal development to be successful, it always requires the interaction of genes, environment, epigenetic processes, and interactions between everything that is used to ‘construct’ the organism and the traits it has. Thus “it makes no sense to ask if a particular trait is genetic or environmental in origin. Understanding how a trait develops is not a matter of finding out whether a particular gene or a particular environment causes the trait; rather, it is a matter of understanding how the various resources available in the production of the trait interact over time” (Kaplan, 2006).
Lastly, I will shortly comment on Sesardic’s (2005: chapter 2) critiques on developmental systems theorists and their critique of heritability and the concept of interactionism. Sesardic argues in the chapter that interaction between genes and environment, nature and nurture, does not undermine heritability estimates (the nature and nurture partition). Philosopher of science Helen Longino argues in her book Studying Human Behavior (2013):
By framing the debate in terms of nature versus nurture and as though one of these must be correct, Sesardic is committed to both downplaying the possible contributions of environmentally oriented research and to relying on a highly dubious (at any rate, nonmethodological) empirical claim.
In sum, the “interactionist fallacy” (coined by Gottfredson) is not a ‘fallacy’ (error in reasoning) at all. For, as Oyama writes in Evolution’s Eye: A Systems View of the Biology-Culture Divide “A not uncommon reaction to DST is, ‘‘That’s completely crazy, and besides, I already knew it” (pg 195). This is exactly what Gottfredson (2009) states, that she “already knew” that there is an interaction between nature and nurture; but she goes on to deny arguments from Oyama, Griffiths, Stotz, Moore, and others on the uselessness of heritability estimates along with the claim that nature and nurture cannot be neatly partitioned into percentages as they are constantly interacting. Causal parity between genes and other developmental resources, too, upends the claim that heritability estimates for any trait make sense (not least for how heritability estimates are gleaned for humans—mostly twin, family, and adoption studies). Developmental denialism—what Gottfredson and others often engage in—runs rampant in the “behavioral genetic” sphere; and Oyama, Griffiths, Stotz, and others show how we should not deny development and we should discard with these estimates for human traits.
Heritability estimates imply that there is a “nature vs nurture” when it is “nature and nurture” which are constantly interacting—and, due to this, we should discard with these estimates due to the interaction of numerous developmental resources; it does not make sense to partition an interacting, self-organizing developmental system. Claims from behavior geneticists—that genes and environment can be separated—are clearly false.
DNA—Blueprint and Fortune Teller?
2500 words
What would you think if you heard about a new fortune-telling device that is touted to predict psychological traits like depression, schizophrenia and school achievement? What’s more, it can tell your fortune from the moment of your birth, it is completely reliable and unbiased — and it only costs £100.
This might sound like yet another pop-psychology claim about gimmicks that will change your life, but this one is in fact based on the best science of our times. The fortune teller is DNA. The ability of DNA to understand who we are, and predict who we will become has emerged in the last three years, thanks to the rise of personal genomics. We will see how the DNA revolution has made DNA personal by giving us the power to predict our psychological strengths and weaknesses from birth. This is a game-changer as it has far-reaching implications for psychology, for society and for each and every one of us.
This DNA fortune teller is the culmination of a century of genetic research investigating what makes us who we are. When psychology emerged as a science in the early twentieth century, it focused on environmental causes of behavior. Environmentalism — the view that we are what we learn — dominated psychology for decades. From Freud onwards, the family environment, or nurture, was assumed to be the key factor in determining who we are. (Plomin, 2018: 6, my emphasis)
The main premise of Plomin’s 2018 book Blueprint is that DNA is a fortune teller while personal genomics is a fortune-telling device. The fortune-telling device Plomin most discusses in the book is polygenic scores (PGS). PGSs are gleaned from GWA studies; SNP genotypes are then added up with scores of 0, 1, and 2. Then, the individual gets their PGS for trait T. Plomin’s claim—that DNA is a fortune teller—though, falls since DNA is not a blueprint—which is where the claim that “DNA is a fortune teller” is derived.
It’s funny that Plomin calls the measure “unbiased”, (he is talking about DNA, which is in effect “unbiased”), but PGS are anything BUT unbiased. For example, most GWAS/PGS are derived from European populations. But, for example, there are “biases and inaccuracies of polygenic risk scores (PRS) when predicting disease risk in individuals from populations other than those used in their derivation” (De La Vega and Bustamante, 2018). (PRSs are derived from statistical gene associations using GWAS; Janssens and Joyner, 2019.) Europeans make up more than 80 percent of GWAS studies. This is why, due to the large amount of GWASs on European populations, that “prediction accuracy [is] reduced by approximately 2- to 5-fold in East Asian and African American populations, respectively” (Martin et al, 2018). See for example Figure 1 from Martin et al (2018):
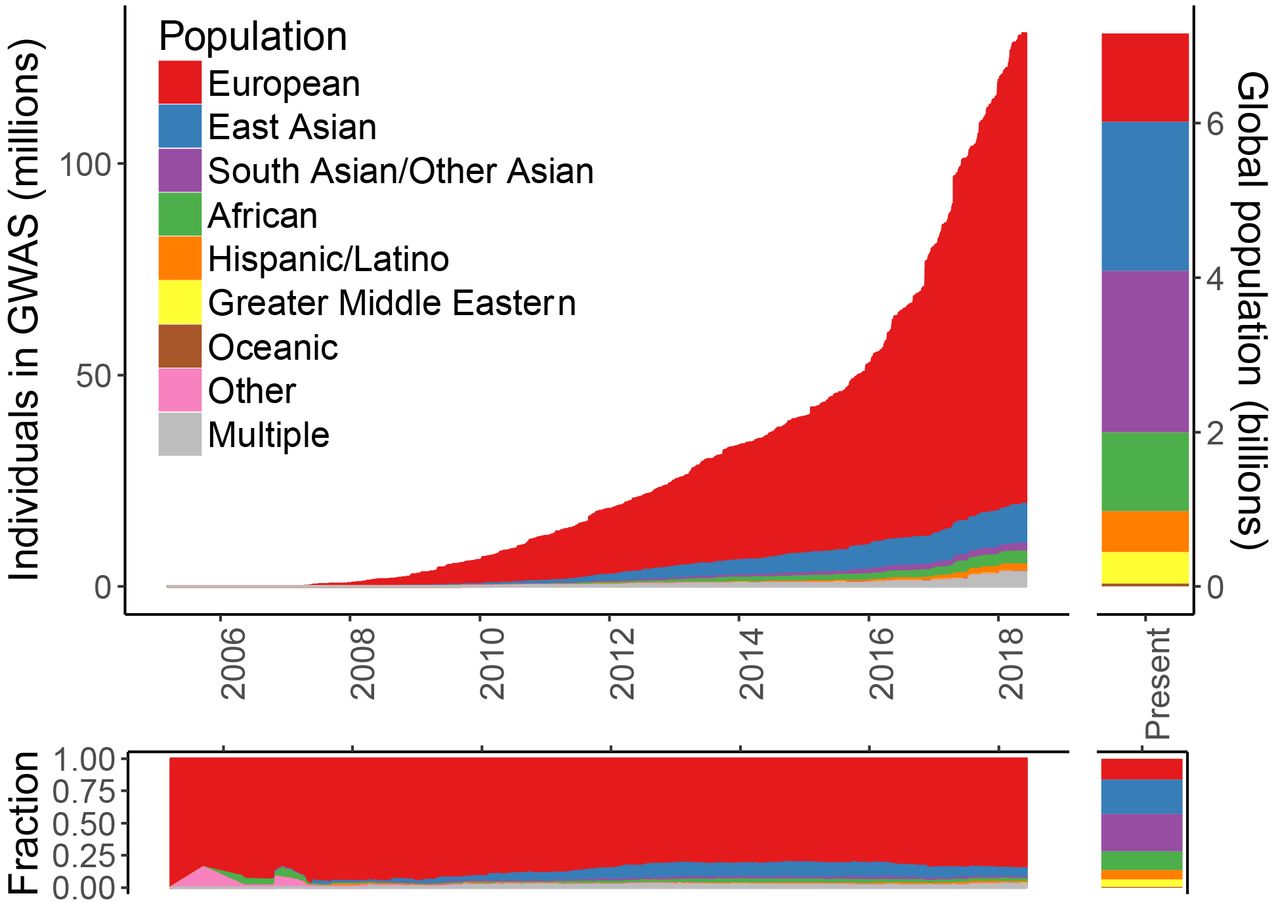
With the huge number of GWAS studies done on European populations, these scores cannot be used on non-European populations for ‘prediction’—even disregarding the other problems with PGS/GWAS.
By studying genetically informative cases like twins and adoptees, behavioural geneticists discovered some of the biggest findings in psychology because, for the first time, nature and nurture could be disentangled.
[…]
… DNA differences inherited from our parents at the moment of conception are the consistent, lifelong source of psychological individuality, the blueprint that makes us who we are. A blueprint is a plan. … A blueprint isn’t all that matters but it matters more than everything else put together in terms of the stable psychological traits that make us who we are. (Plomin, 2018: 6-8, my emphasis)
Nevermind the slew of problems with twin and adoption studies (Joseph, 2014; Joseph et al, 2015; Richardson, 2017a). I also refuted the notion that “A blueprint is a plan” last year, quoting numerous developmental systems theorists. The main thrust of Plomin’s book—that DNA is a blueprint and therefore can be seen as a fortune teller using the fortune-telling device to tell the fortunes of the people’s whose DNA are analyzed—is false, as DNA does not work how it does in Plomin’s mind.
These big findings were based on twin and adoption studies that indirectly assessed genetic impact. Twenty years ago the DNA revolution began with the sequencing of the human genome, which identified each of the 3 billion steps in the double helix of DNA. We are the same as every other human being for more than 99 percent of these DNA steps, which is the blueprint for human nature. The less than 1 per cent of difference of these DNA steps that differ between us is what makes us who we are as individuals — our mental illnesses, our personalities and our mental abilities. These inherited DNA differences are the blueprint for our individuality …
[DNA predictors] are unique in psychology because they do not change during our lives. This means that they can foretell our futures from our birth.
[…]
The applications and implications of DNA predictors will be controversial. Although we will examine some of these concerns, I am unabashedly a cheerleader for these changes. (Plomin, 2018: 8-10, my emphasis)
This quote further shows Plomin’s “blueprint” for the rest of his book—DNA can “foretell our futures from our birth”—and how it affects his conclusions gleaned from his work that he mostly discusses in his book. Yes, all scientists are biased (as Stephen Jay Gould noted), but Plomin outright claimed to be an unabashed cheerleader for his work. Plomin’s self-admission for being an “unabashed cheerleader”, though, does explain some of the conclusions he makes in Blueprint.
However, the problem with the mantra ‘nature and nurture’ is that it runs the risk of sliding back into the mistaken view that the effects of genes and environment cannot be disentangled.
[…]
Our future is DNA. (Plomin, 2018: 11-12)
The problem with the mantra “nature and nurture” is not that it “runs the risk of sliding back into the mistaken view that the effects of genes and environment cannot be disentangled”—though that is one problem. The problem is how Plomin assumes how DNA works. That DNA can be disentangled from the environment presumes that DNA is environment-independent. But as Moore shows in his book The Dependent Gene—and as Schneider (2007) shows—“the very concept of a gene requires the environment“. Moore notes that “The common belief that genes contain context-independent “information”—and so are analogous to “blueprints” or “recipes”—is simply false” (quoted in Schneider, 2007). Moore showed in The Dependent Gene that twin studies are flawed, as have numerous other authors.
Lewkowicz (2012) argues that “genes are embedded within organisms which, in turn, are embedded in external environments. As a result, even though genes are a critical part of developmental systems, they are only one part of such systems where interactions occur at all levels of organization during both ontogeny and phylogeny.” Plomin—although he does not explicitly state it—is a genetic reductionist. This type of thinking can be traced back, most popularly, to Richard Dawkins’ 1976 book The Selfish Gene. The genetic reductionists can, and do, make the claim that organisms can be reduced to their genes, while developmental systems theorists claim that holism, and not reductionism, better explains organismal development.
The main thrust of Plomin’s Blueprint rests on (1) GWA studies and (2) PGSs/PRSs derived from the GWA studies. Ken Richardson (2017b) has shown that “some cryptic but functionally irrelevant genetic stratification in human populations, which, quite likely, will covary with social stratification or social class.” Richardson’s (2017b) argument is simple: Societies are genetically stratified; social stratification maintains genetic stratification; social stratification creates—and maintains—cognitive differentiation; “cognitive” tests reflect prior social stratification. This “cryptic but functionally irrelevant genetic stratification in human populations” is what GWA studies pick up. Richardson and Jones (2019) extend the argument and argue that spurious correlations can arise from genetic population structure that GWA studies cannot account for—even though GWA study authors claim that this population stratification is accounted for, social class is defined solely on the basis of SES (socioeconomic status) and therefore, does not capture all of what “social class” itself captures (Richardson, 2002: 298-299).
Plomin also heavily relies on the results of twin and adoption studies—a lot of it being his own work—to attempt to buttress his arguments. However, as Moore and Shenk (2016) show—and as I have summarized in Behavior Genetics and the Fallacy of Nature vs Nurture—heritability estimates for humans are highly flawed since there cannot be a fully controlled environment. Moore and Shenk (2016: 6) write:
Heritability statistics do remain useful in some limited circumstances, including selective breeding programs in which developmental environments can be strictly controlled. But in environments that are not controlled, these statistics do not tell us much. In light of this, numerous theorists have concluded that ‘the term “heritability,” which carries a strong conviction or connotation of something “[in]heritable” in the everyday sense, is no longer suitable for use in human genetics, and its use should be discontinued.’ 31 Reviewing the evidence, we come to the same conclusion.
Heritability estimates assume that nature (genes) can be separated from nurture (environment), but “the very concept of a gene requires the environment” (Schneider, 2007) so it seems that attempting to partition genetic and environmental causation of any trait T is a fool’s—reductionist—errand. If the concept of gene depends on and requires the environment, then how does it make any sense to attempt to partition one from the other if they need each other?
Let’s face it: Plomin, in this book Blueprint is speaking like a biological reductionist, though he may deny the claim. The claims from those who push PRS and how it can be used for precision medicine are unfounded, as there are numerous problems with the concept. Precision medicine and personalized medicine are similar concepts, though Joyner and Paneth (2015) are skeptical of its use and have seven questions for personalized medicine. Furthermore, Joyner, Boros and Fink (2018) argue that “redundant and degenerate mechanisms operating at the physiological level limit both the general utility of this assumption and the specific utility of the precision medicine narrative.” Joyner (2015: 5) also argues that “Neo-Darwinism has failed clinical medicine. By adopting a broader perspective, systems biology does not have to.”
Janssens and Joyner (2019) write that “Most [SNP] hits have no demonstrated mechanistic linkage to the biological property of interest.” Researchers can show correlations between disease phenotypes and genes, but they cannot show causation—which would be mechanistic relations between the proposed genes and the disease phenotype. Though, as Kampourakis (2017: 19), genes do not cause diseases on their own, they only contribute to its variation.
Edit: Take also this quote from Plomin and Stumm (2018) (quoted by Turkheimer):
GPS are unique predictors in the behavioural sciences. They are an exception to the rule that correlations do not imply causation in the sense that there can be no backward causation when GPS are correlated with traits. That is, nothing in our brains, behaviour or environment changes inherited differences in DNA sequence. A related advantage of GPS as predictors is that they are exceptionally stable throughout the life span because they index inherited differences in DNA sequence. Although mutations can accrue in the cells used to obtain DNA, like any cells in the body these mutations would not be expected to change systematically the thousands of inherited SNPs that contribute to a GPS.
Turkheimer goes on to say that this (false) assumption by Plomin and Stumm (2018) assumes that there is no top-down causation—i.e., that phenotypes don’t cause genes, or there is no causation from the top to the bottom. (See the special issue of Interface Focus for a slew of articles on top-down causation.) Downward causation exists in biological systems (Noble, 2012, 2017), as does top-down. The very claim that “nothing in our brains, behaviour or environment changes inherited differences in DNA sequence” is ridiculous! This is something that, of course, Plomin did not discuss in Blueprint. But in a book that, supposedly, shows “how DNA makes us who we are”, why not discuss epigenetics? Plomin is confused, because DNA methylation impacts behavior and behavior impacts DNA methylation (Lerner and Overton, 2017: 114). Lerner and Overtone (2017: 145) write that:
… it should no longer be possible for any scientist to undertake the procedure of splitting of nature and nurture and, through reductionist procedures, come to conclusions that the one or the other plays a more important role in behavior and development.
Plomin’s reductionist takes, therefore again, fail. Plomin’s “reluctance” to discuss “tangential topics” to “inherited DNA differences” included epigenetics (Plomin, 2018: 12). But it seems that his “reluctance” to discuss epigenetics was a downfall in his book as epigenetic mechanisms can and do make a difference to “inherited DNA differences” (see for example, Baedke, 2018, Above the Gene, Beyond Biology: Toward a Philosophy of Epigenetics and Meloni, 2019, Impressionable Biologies: From the Archaeology of Plasticity to the Sociology of Epigenetics see also Meloni, 2018). The genome can and does “react” to what occurs to the organism in the environment, so it is false that “nothing in our brains, behaviour or environment changes inherited differences in DNA sequence” (Plomin and Stumm, 2018), since our behavior and actions can and do methylate our DNA (Meloni, 2014) which falsifies Plomin’s claim and which is why he should have discussed epigenetics in Blueprint. End Edit
Conclusion
So the main premise of Plomin’s Blueprint is his two claims: (1) that DNA is a fortune teller and (2) that personal genomics is a fortune-telling device. He draws these big claims from PGS/PRS studies. However, over 80 percent of GWA studies have been done on European populations. And, knowing that we cannot use these datasets on other, non-European datasets, greatly hampers the uses of PGS/PRS in other populations—although the PGS/PRS are not that useful in and of itself for European populations. Plomin’s whole book is a reductionist screed—“Sure, other factors matter, but DNA matters more” is one of his main claims. Though, a priori, since there is no privileged level of causation, one cannot privilege DNA over any other developmental variables (Noble, 2012). To understand disease, we must understand the whole system and how when one part of the system becomes dysfunctional how it affects other parts of the system and how it runs. The PGS/PRS hunts are reductionist in nature, and the only answer to these reductionist paradigms are new paradigms from systems biology—one of holism.
Plomin’s assertions in his book are gleaned from highly confounded GWA studies. Plomin also assumes that we can disentangle nature and nurture—like all reductionists. Nature and nurture interact—without genes, there would be an environment, but without an environment, there would be no genes as gene expression is predicated on the environment and what occurs in it. So Plomin’s reductionist claim that “Our future is DNA” is false—our future is studying the interactive developmental system, not reducing it to a sum of its parts. Holistic biology—systems biology—beats reductionist biology—the Neo-Darwinian Modern Synthesis.
DNA is not a blueprint nor is it a fortune teller and personal genomics is not a fortune-telling device. The claim that DNA is a blueprint/fortune teller and personal genomics is a fortune-telling device come from Plomin and are derived from highly flawed GWA studies and, further, PGS/PRS. Therefore Plomin’s claim that DNA is a blueprint/fortune teller and personal genomics is a fortune-telling device are false.
(Also read Erick Turkheimer’s 2019 review of Plomin’s book The Social Science Blues, along with Steve Pitteli’s review Biogenetic Overreach for an overview and critiques of Plomin’s ideas. And read Ken Richardson’s article It’s the End of the Gene As We Know It for a critique of the concept of the gene.)
Twin Studies, Adoption Studies, and Fallacious Reasoning
2350 words
Twin and adoption studies have been used for decades on the basis that genetic and environmental causes of traits and their variation in the population could be easily partitioned by two ways: one way is to adopt twins into separate environments, the other to study reared-together or reared-apart twins. Both methods rest on a large number of (invalid) assumptions. These assumptions are highly flawed and there is no evidential basis to believe these assumptions, since the assumptions have been violated which invalidates said assumptions.
Plomin et al write (2013) write: For nearly a century, twin and adoption studies have yielded substantial estimates of heritability for cognitive abilities.
But the validity of the “substantial estimates of heritability for cognitive abilities” is strongly questioned due to unverified (and false) assumptions that these researchers make.
Adoption studies
The problem with adoption studies are numerous, not least: restricted range of adoptive families; selective placement; late separation; parent-child attachment disturbance; problems with the tests (on personality, ‘IQ’); the non-representativeness of adoptees compared to non-adoptees; and the reliability of the characteristic in question.
In selective placement, the authorities attempt to place children in homes close to their biological parents. They gage how “intelligent” they believe they are (on the basis of parental SES and the child’s parent’s perceived ‘intelligence’), thusly this is a pretty huge confound for adoption studies.
According to adoption researcher Harry Munsinger, a “possible source of bias in adoption studies is selective placement of adopted children in adopting homes that are similar to their biological parents’ social and educational backgrounds.” He recognized that “‘fitting the home to the child’ has been the standard practice in most adoption agencies, and this selective placement can confound genetic endowment with environmental influence to invalidate the basic logic of an adoptive study (Munsinger, 1975, p. 627). Clearly, agency policies of “fitting the home to the child” are a far cry from random placement of adoptees into a wide range of adoptive homes. (Joseph, 2015: 30-1)
Richardson and Norgate (2005) argue that simple additive effects for both genetic and environmental effects are false; that IQ is not a quantitative trait; while other interactive effects could explain the IQ correlation.
1) Assignment is nonrandom. 2) They look for adoptive homes that reflect the social class of the biological mother. 3) This range restriction reduces the correlation estimates between adopted children and adopted parents. 4) Adoptive mothers come from a narrow social class. 5) Their average age at testing will be closet to their biological parents than adopted parents. 6) They experience the womb of their mothers. 7) Stress in the womb can alter gene expression. 8) Adoptive parents are given information about the birth family which may bias their treatment. 9) Biological mothers and adopted children show reduced self-esteem and are more vulnerable to changing environments which means they basically share environment. 10) Conscious or unconscious aspects of family treatment may make adopted children different from other adopted family members. 11) Adopted children also look more like their biological parents than their adoptive parents which means they’ll be treated accordingly.
Twin studies
Personally, my favorite thing to discuss. Twin studies rest on the erroneous assumption that DZ and MZ environments are equal; that they get treated equally the same. This is false, MZ twins get treated more similarly than DZ twins, which twin researchers have conceded decades ago. But in order to save their field, they attempt to use circular argumentation, known as Argument A. Argument A states that MZTs (monozygotic twins reared together) are more genetically similar than DZTs (dizygotic twins reared together) and thusly this causes greater behavioral similarity. But this is based on circular reasoning: the researchers already implicitly assumed that genes played a role in their premise and, not surprisingly, in their conclusion genes are the cause for the similarities of the MZTs. So Argument A is used, twin researchers circularly assume that MZTs greater behavioral similarity is due to genetic similarity, while their argument that genetic factors explain the greater behavioral similarity of MZTs is a premise and conclusion of their argument. “X is true because Y is true; Y is true because X is true.” (Also see Joseph et al, 2015.)
We have seen that circular reasoning is “empty reasoning in which the conclusion rests on an assumption whose validity is dependent on the conclusion” (Reber, 1985, p. 123). … A circular argument consists of “using as evidence a fact which is authenticated by the very conclusion it supports,” which “gives us two unknowns so busy chasing each other’s tails that neither has time to attach itself to reality” (Pirie, 2006, p. 27) (Joseph, 2016: 164).
Even if Argument A is accepted, the causes of behavioral similarities between MZ/DZ twins could still come down to environment. Think of any type of condition that is environmentally caused but is due to people liking what causes the condition. There are no “genes for” that condition, but their liking the thing that caused the condition caused an environmental difference.
Argument B also exists. Those that use Argument B also concede that MZs experience more similar environments, but then argue that in order to show that twin studies, and the EEA, are false, critics must show that MZT and DZT environments differ in the aspects that are relevant to the behavior in question (IQ, schizophrenia, etc).
An example of an Argument B environmental factor relevant to a characteristic or disorder is the relationship between exposure to trauma and post-traumatic stress disorder (PTSD). Because trauma exposure is (by definition) an environmental factor known to contribute to the development of PTSD, a finding that MZT pairs are more similarly exposed to trauma than DZT pairs means that MZT pairs experience more similar “trait-relevant” environments than DZTs. Many twin researchers using Argument B would conclude that the EEA is violated in this case. (Joseph, 2016: 165)
So twin researchers need to rule out and identify “trait-relevant factors” which contribute to the cause of said trait, along with experiencing more similar environments, invalidates genetic interpretations made using Argument B. But Argument A renders Argument B irrelevant because even if critics can show that MZTs experience more similar “trait-relevant environments”, they could still argue that the twin method is valid by stating that (in Argument A fashion) MZTs create and elicit more similar trait-relevant environments.
One more problem with Argument A is that it shows that twins behave accordingly to “inherited environment-creating blueprint” (Joseph, 2016: 164) but at the same time shows that parents and other adults are easily able to change their behaviors to match that of the behaviors that the twins show, which in effect, allows them to “create” or “elicit” their own environments. But the adults’ “environment-creating behavior and personality” should be way more unchangeable than the twins’ since along with the presumed genetic similarity, adults have “experienced decades of behavior-molding peer, family, religious, and other socialization influences” (Joseph, 2016: 165).
Whether or not circular arguments are “useful” or not has been debated in the philosophical literature for some time (Hahn, Oaksford, and Corner, 2005). However, assuming, in your premise, that your conclusion is valid is circular and therefore While circular arguments are deductively valid, “it falls short of the ultimate goal of argumentative discourse: Whatever evaluation is attached to the premise is transmitted to the conclusion, where it remains the same; no increase in degree of belief takes place” (Hahn, 2011: 173).
However, Hahn (2011: 180) concludes that “the existence of benign circularities makes clear that merely labeling something as circular is not enough to dismiss it; an argument for why the thing in question is bad still needs to be made.” This can be simply shown: The premise that twin researchers use (that genes cause similar environments to be constructed) is in their conclusion. They state in their premise that MZT behavioral similarity is due to greater MZT genetic similarity in comparison to DZTs (100 vs. 50 percent). Then, in the conclusion, they re-state that the behavioral similarities of MZTs is due to their genetic similarity compared to DZTs (100 vs. 50 percent). Thus, a convincing argument for conclusion C (that genetic similarity explains MZT behavioral similarity) cannot rest on the assumption that conclusion C is correct. Thus, Argument A is fallacious due to its circularity.
What causes MZT behavioral similarities is their more similar environment: they get treated the same by peers and parents, and have higher rates of identity confusion and had a closer emotional bond compared to DZTs. The twin method is based on the (erroneous) assumption that MZT and DZT pairs experience roughly equal environments, which twin researchers conceded was false decades ago.
Richardson and Norgate (2005: 347) conclude (emphasis mine):
We have shown, first, that the EEA may not hold, and that well-demonstrated treatment effects can, therefore, explain part of the classic MZ–DZ differences. Using published correlations, we have also shown how sociocognitive interactions, in which DZ twins strive for a relative ‘apartness’, could further depress DZ correlations, thereby possibly explaining another part of the differences. We conclude that further conclusions about genetic or environmental sources of variance from MZ–DZ twin data should include thorough attempts to validate the EEA with the hope that these interactions and their implications will be more thoroughly understood.
Of course, even if twin studies were valid and the EEA was true/ the auxiliary arguments used were true, this would still not mean that heritability estimates would be of any use to humans, since we cannot control environments as we do in animal breeding studies (Schonemann, 1997; Moore and Shenk, 2016). I have chronicled how 1) the EEA is false and how flawed twin studies are; 2) how flawed heritability estimates are; 3) how heritability does not (and cannot) show causation; and 4) the genetic reductionist model that behavioral geneticists rely on is flawed (Lerner and Overton, 2017).
So we can (1) accept the EEA, that the greater behavioral resemblance indicates the importance of genetic factors underlying most human behavioral differences and behavioral disorders or we can (2) reject the EEA and state that the greater behavioral resemblance is due to nongenetic (environmental) factors, which means that all genetic interpretations of MZT/DZT studies must be rejected. Thus, using (2), we can infer that all twin studies measure is similarity of the environment of DZTs, and it is, in fact, not measuring genetic factors. Accepting explanation 2 does not mean that “twin studies overestimate heritability, or that researchers should assess the EEA on a study-by-study basis, but instead indicates that the twin method is no more able than a family study to disentangle the potential influences of genes and environment” (Joseph, 2016: 181).
What it does mean, however, is that we can, logically, discard all past, future, and present MZT and DZT comparisons and these genetic interpretations must be outright rejected, due to the falsity of the EEA and the fallaciousness of the auxiliary arguments made in order to save the EEA and the twin method overall.
There are further problems with twin studies and heritability estimates. Epigenetic supersimilarity (ESS) also confounds the relationship. Due to the existence of ESS “human MZ twins clearly cannot be viewed as the epigenetic equivalent of isogenic inbred mice, which originate from separate zygotes. To the extent that epigenetic variation at ESS loci influences human phenotype, as our data indicate, the existence of ESS establishes a link between early embryonic epigenetic development and adult disease and may call into question heritability estimates based on twin studies” (Van Baak et al, 2018). In other words, ESS is an unrecognized phenomenon that contributes to the phenotypic similarity of MZs, which calls into question the usefulness of heritability studies using twins. The uterine environment has been noted to be a confound by numerous authors (Devlin, Daniels, and Roeder, 1997; Charney, 2012; Ho, 2013; Moore and Shenk, 2016).
Conclusion
Adoption studies fall prey to numerous pitfalls, most importantly, that children are adopted into similar homes compared to their birth parents, which restricts the range of environments for adoptees. Adoption placement is also non-random, the children are placed into homes that are similar to their biological parents. Due to these confounds (and a whole slew of other invalidating problems), adoption studies cannot be said to show genetic causation, nor can they separate genetic from environmental factors.
Twin studies suffer from the biggest flaw of all: the falsity of the EEA. Since the EEA is false—which has been recognized by both critics and supporters of the assumption—the supporters of the assumption have attempted to redefine the EEA in two ways: (1) that MZTs experience more similar environments due to genetic similarity (Argument A) and (2) that it is not whether MZTs experience more similar environments, but whether or not they share more similar trait-relevant environments. Thus, unless these twin researchers are able to identify trait-relevant factors that contribute to the trait in question, we must conclude that (along with the admission from twin researchers that the EEA is false; that MZTs experience more similar environments than DZTs) genetic interpretations made using Argument B are thusly invalidated. Fallacious reasoning (“X causes Y; Y causes X) does not help any twin argument. Because their conclusion is already implicitly assumed in their premise.
The existence of ESS (epigenetic supersimilarity) further shows how invalid the twin method truly is, because the confounding starts in the womb. Attempts can be made (however bad) to control for shared environment by adopting different twins into different homes, but they still shared a uterine environment which means they shared an environment, which means it is a confound and it cannot be controlled for (Charney, 2012).
Adoption and twin studies are highly flawed. Like family studies, twin studies are no more able to disentangle genetic from environmental effects than a family study, and thus twin studies cannot separate genes from environment. Last, and surely not least, it is fallacious to assume that genes can be separated so neatly into “heritability estimates” as I have noted in the past. Heritability estimates cannot show genetic causation, nor can it show how malleable a trait is. They’re just (due to how we measure) flawed measures that we cannot fully control so we must make a number of (false) assumptions that then invalidate the whole paradigm. The EEA is false, all auxiliary arguments made to save the EEA are fallacious; adoption studies are hugely confounded; twin studies are confounded due to numerous reasons, most importantly the uterine environment (Van Baak et al, 2018).
Find the Genes: Testosterone Version
1600 words
Testosterone has a similar heritability to IQ (between .4 and .6; Harris, Vernon, and Boomsma, 1998; Travison et al, 2014). To most, this would imply a significant effect of genes on the production of testosterone and therefore we should find a lot of SNPs that affect the production of testosterone. However, testosterone production is much more complicated than that. In this article, I will talk about testosterone production and discuss two studies which purport to show a few SNPs associated with testosterone. Now, this doesn’t mean that the SNPs cause high/low testosterone, just that they were associated. I will then speak briefly on the ‘IQ SNPs’ and compare it to ‘testosterone SNPs’.
Testosterone SNPs?
Complex traits are ‘controlled’ by many genes and environmental factors (Garland Jr., Zhao, and Saltzman, 2016). Testosterone is a complex trait, so along with the heritability of testosterone being .4 to .6, there must be many genes of small effect that influence testosterone, just like they supposedly do for IQ. This is obviously wrong for testosterone, which I will explain below.
Back in 2011 it was reported that genetic markers were discovered ‘for’ testosterone, estrogen, and SHGB production, while showing that genetic variants in the SHGB locus, located on the X chromosome, were associated with substantial testosterone variation and increased the risk of low testosterone (important to keep in mind) (Ohlsson et al, 2011). The study was done since low testosterone is linked to numerous maladies. Low testosterone is related to cardiovascular risk (Maggio and Basaria, 2009), insulin sensitivity (Pitteloud et al, 2005; Grossman et al, 2008), metabolic syndrome (Salam, Kshetrimayum, and Keisam, 2012; Tsuijimora et al, 2013), heart attack (Daka et al, 2015), elevated risk of dementia in older men (Carcaillon et al, 2014), muscle loss (Yuki et al, 2013), and stroke and ischemic attack (Yeap et al, 2009). So this is a very important study to understand the genetic determinants of low serum testosterone.
Ohlsson et al (2011) conducted a meta-analysis of GWASs, using a sample of 14,429 ‘Caucasian’ men. To be brief, they discovered two SNPs associated with testosterone by performing a GWAS of serum testosterone concentrations on 2 million SNPs on over 8,000 ‘Caucasians’. The strongest associated SNP discovered was rs12150660 was associated with low testosterone in this analysis, as well as in a study of Han Chinese, but it is rare along with rs5934505 being associated with an increased risk of low testosterone(Chen et al, 2016). Chen et al (2016) also caution that their results need replication (but I will show that it is meaningless due to how testosterone is produced in the body).
Ohlsson et al (2011) also found the same associations with the same two SNPs, along with rs6258 which affect how testosterone binds to SHGB. Ohlsson et al (2011) also validated their results: “To validate the independence of these two SNPs, conditional meta-analysis of the discovery cohorts including both rs12150660 and rs6258 in an additive genetic linear model adjusted for covariates was calculated.” Both SNPs were independently associated with low serum testosterone in men (less than 300ng/dl which is in the lower range of the new testosterone guidelines that just went into effect back in July). Men who had 3 or more of these SNPs were 6.5 times more likely to have lower testosterone.
Ohlsson et al (2011) conclude that they discovered genetic variants in the SHGB locus and X chromosome that significantly affect serum testosterone production in males (noting that it’s only on ‘Caucasians’ so this cannot be extrapolated to other races). It’s worth noting that, as can be seen, these SNPs are not really associated with variation in the normal range, but near the lower end of the normal range in which people would then need to seek medical help for a possible condition they may have.
In infant males, no SNPs were significantly associated with salivary testosterone levels, and the same was seen for infant females. Individual variation in salivary testosterone levels during mini-puberty (Kurtoglu and Bastug, 2014) were explained by environmental factors, not SNPs (Xia et al, 2014). They also replicated Carmaschi et al (2010) who also showed that environmental factors influence testosterone more than genetic factors in infancy. There is a direct correlation between salivary testosterone levels and free serum testosterone (Wang et al, 1981; Johnson, Joplin, and Burin, 1987), so free serum testosterone was indirectly tested.
This is interesting because, as I’ve noted here numerous times, testosterone is indirectly controlled by DNA, and it can be raised or lowered due to numerous environmental variables (Mazur and Booth, 1998; Mazur, 2016), such as marriage (Gray et al, 2002; Burnham et al, 2003; Gray, 2011; Pollet, Cobey, and van der Meij, 2013; Farrelly et al, 2015; Holmboe et al, 2017), having children (Gray et al, 2002; Gray et al, 2006; Gettler et al, 2011); to obesity (Palmer et al, 2012; Mazur et al, 2013; Fui, Dupuis, and Grossman, 2014; Jayaraman, Lent-Schochet, and Pike, 2014; Saxbe et al, 2017) smoking is not clearly related to testosterone (Zhao et al, 2016), and high-carb diets decrease testosterone (Silva, 2014). Though, most testosterone decline can be ameliorated with environmental interventions (Shi et al, 2013), it’s not a foregone conclusion that testosterone will sharply decrease around age 25-30.
Studies on ‘testosterone genes’ only show associations, not causes, genes don’t directly cause testosterone production, it is indirectly controlled by DNA, as I will explain below. These studies on the numerous environmental variables that decrease testosterone is proof enough of the huge effects of environment on testosterone production and synthesis.
How testosterone is produced in the body
There are five simple steps to testosterone production: 1) DNA codes for mRNA; 2) mRNA codes for the synthesis of an enzyme in the cytoplasm; 3) luteinizing hormone stimulates the production of another messenger in the cell when testosterone is needed; 4) this second messenger activates the enzyme; 5) the enzyme then converts cholesterol to testosterone (Leydig cells produce testosterone in the presence of luteinizing hormone) (Saladin, 2010: 137). Testosterone is a steroid and so there are no ‘genes for’ testosterone.
Cells in the testes enzymatically convert cholesterol into the steroid hormone testosterone. Quoting Saladin (2010: 137):
But to make it [testosterone], a cell of the testis takes in cholesterol and enzymatically converts it to testosterone. This can occur only if the genes for the enzymes are active. Yet a further implication of this is that genes may greatly affect such complex outcomes as behavior, since testosterone strongly influences such behaviors as aggression and sex drive. [RR: Most may know that I strongly disagree with the fact that testosterone *causes* aggression, see Archer, Graham-Kevan and Davies, 2005.] In short, DNA codes only for RNA and protein synthesis, yet it indirectly controls the synthesis of a much wider range of substances concerned with all aspects of anatomy, physiology, and behavior.

(Figure from Saladin (2010: 137; Anatomy and Physiology: The Unity of Form and Function)
Genes only code for RNA and protein synthesis, and thusly, genes do not *cause* testosterone production. This is a misconception most people have; if it’s a human trait, then it must be controlled by genes, ultimately, not proximately as can be seen, and is already known in biology. Genes, on their own, are not causes but passive templates (Noble, 2008; Noble, 2011; Krimsky, 2013; Noble, 2013; Also read Exploring Genetic Causation in Biology). This is something that people need to understand; genes on their own do nothing until they are activated by the system.
What does this have to do with ‘IQ genes’?
My logic here is very simple: 1) Testosterone has the same heritability range as IQ. 2) One would assume—like is done with IQ—that since testosterone is a complex trait that it must be controlled by ‘many genes of small effect’. 3) Therefore, since I showed that there are no ‘genes for’ testosterone and only ‘associations’ (which could most probably be mediated by environmental interventions) with low testosterone, may the same hold true for ‘IQ genes/SNPS’? These testosterone SNPs I talked about from Ohlsson et al (2011) were associated with low testosterone. These ‘IQ SNP’ studies (Davies et al, 2017; Hill et al, 2017; Savage et al, 2017) are the same—except we have an actual idea of how testosterone is produced in the body, we know that DNA is indirectly controlling its production, and, most importantly, there is/are no ‘gene[s] for’ testosterone.
Conclusion
Testosterone has the same heritability range as IQ, is a complex trait like IQ, but, unlike how IQ is purported to be, it [testosterone] is not controlled by genes; only indirectly. My reasoning for using this example is simple: something has a moderate to high heritability, and so most would assume that ‘numerous genes of small effect’ would have an influence on testosterone production. This, as I have shown, is false. It’s also important to note that Ohlsson et al (2011) showed associated SNPs in regards to low testosterone—not testosterone levels in the normal range. Of course, only when physiological values are outside of the normal range will we notice any difference between men, and only then will we find—however small—genetic differences between men with normal and low levels of testosterone (I wouldn’t be surprised if lifestyle factors explained the lower testosterone, but we’ll never know that in regards to this study).
Testosterone production is a real, measurable physiologic process, as is the hormone itself; which is not unlike the so-called physiologic process that ‘g’ is supposed to be, which does not mimic any known physiologic process in the body, which is covered with unscientific metaphors like ‘power’ and ‘energy’ and so on. This example, in my opinion, is important for this debate. Sure, Ohlsson et al (2011) found a few SNPs associated with low testosterone. That’s besides the point. They are only associated with low testosterone; they do not cause low testosterone. So, I assert, these so-called associated SNPs do not cause differences in IQ test scores; just because they’re ‘associated’ doesn’t mean they ’cause’ the differences in the trait in question. (See Noble, 2008; Noble, 2011; Krimsky, 2013; Noble, 2013.) The testosterone analogy that I made here buttresses my point due to the similarities (it is a complex trait with high heritability) with IQ.
Heritability, the Grandeur of Life, and My First Linkfest on Human Evolution and IQ
1100 words
Benjamin Steele finally replied to my critique of his ‘strong evidence and argument’ on race, IQ and adoption. He goes on to throw baseless ad hominem attacks as well as appealing to motive (assuming my motivation for being a race realist; assuming that I’m a ‘racist’, whatever that means). When I do address his ‘criticisms’ of my response to him, I will not address his idiotic attacks as they are a waste of time. He does, however, say that I do not understand heritability. I understand that the term ‘heritable’ doesn’t mean ‘genetic’. I understand that heritability is the proportion of phenotypic variance attributed to genetic variance. I do not believe that heritability means a trait is X percent genetic. 80 percent of the variation in the B-W IQ gap is genetic, with 20 percent explained by environmental effects. Note that I’m not claiming that heritable means genetic. All that aside, half of his reply to me is full of idiotic, baseless and untrue accusations which I will not respond to. So, Mr. Steele, if you do decide to reply to my response to you this weekend, please leave the idiocy at the door. Anyway, I will tackle that this weekend. Quick note for Mr. Steele (in case he reads this): if you don’t believe me about the National Crime and Victimization Survey showing that police arrest FEWER blacks than are reported by the NCVS, you can look it up yourself, ya know.
I’m beginning to understand why people become environmentalists. I’ve recently become obsessed with evolution. Not only of Man, but of all of the species in the world. Really thinking about the grandeur of life and evolution and what leads to the grand diversity of life really had me thinking one day. It took billions of years for us to get to the point we did today. So, why should we continue to destroy environments, displacing species and eventually leading them to extinction? I’m not saying that I fully hold this view yet, it’s just been on my mind lately. Once a species is extinct, that’s it, it’s gone forever. So shouldn’t we do all we possibly can to preserve the wonder of life that took so long to get to the point that we did today?
Some interesting articles to read:
Study: IQ of firstborns differ from siblings (This is some nice evidence for Lassek and Gaulin’s theory stating why first-born children have higher IQs than their siblings: they get first dibs on the gluteofemoral fat deposits that are loaded with n-3 fatty acids, aiding in brain size and IQ.)
Why attitude is more important than IQ (Psychologist Carol Dweck states that attitude is more important than IQ and that attitudes come in one of two types: a fixed mindset or a growth mindset. Those with a fixed mindset believe ‘you are who you are’ and nothing can change it while those with a growth mindset believe they can improve with effort. Interesting article, I will find the paper and comment on it when I read it.)
Positively Arguing IQ Determinism And Effect Of Education (Intelligent people search for intellectually stimulating things whereas less intelligent people do not. This, eventually, will lead to the construction of environments based on that genotype.)
A scientist’s new theory: Religion was key to humans’ social evolution (Nicholas Wade pretty much argues the same in his book The Faith Instinct: How Religion Evolved and Why It Endures. It is interesting to note that archaeologists have discovered what looks to be the beginnings of religiousity around 10kya, coinciding with the agrigultural revolution. I will look into this in the future.)
Galápagos giant tortoises show that in evolution, slow and steady gets you places (Interesting read, on tortoise migration)
Will Mars Colonists Evolve Into This New Kind of Human? (Very interesting and I hope to see more articles like this in the future. Of course, due to being a smaller population, evolution will occur faster due to differing selection pressures. Smaller populations incur more mutations at a faster rate than larger populatons. Will our skin turn a reddish tint? Bone density will decline leading to heavier bones. The need for C-sections due to heavier bones will lead to futher brain size increases. This is also going on on Earth at the moment, as I have previously discussed. Of course differences in culture and technology will lead the colonizers down different paths. I hope I am alive to see the first colonies on Mars and the types of long-term effects of the evolution of Man on the Red Planet.)

Evolution debate: Are humans continuing to evolve? (Of course we are)
Did seaweed make us who we are today? (Seaweed has many important vitamins and minerals that are imperative for brain development and growth—most importantly, it has poly-unsaturated fatty acids (PUFAs) and B12. We are only able to acquire these fatty acids through our diet—our body cannot synthesize the fatty acid on its own. This is just growing evidence for how important it is to have a good ratio of n-3 to n-6.)
Desert people evolve to drink water poisoned with deadly arsenic (More evidence for rapid evolution in human populations. AS3MT is known to improve arsonic metabolism in Chile and Argentina. Clearly, those who can handle the water breed/don’t die while those who cannot succumb to the effects of arsenic poisoning. Obviously, over time, this SNP will be selected for more and more while those who cannot metabolize the arsonic do not pass on their genes. This is a great article to show to anti-human-evolution deniers.)
Here Are the Weird Ways Humans Are Evolving Right Now (CRISPR and gene editing, promotion of obesity through environmental factors (our animals have also gotten fatter, probably due to the feed we give them…), autism as an adaptation (though our definition for autism has relaxed in the past decade). Human evolution is ongoing and never stops, even for Africans. I’ve seen some people claim that since they never left the continent that they are ‘behind in evolution’, yet evolution is an ongoing process and never stops, cultural ‘evolution’ (change) leading to more differences.)
‘Goldilocks’ genes that tell the tale of human evolution hold clues to variety of diseases (We really need to start looking at modern-day diseases through an evolutionary perspective, such as obesity, to better understand why these ailments inflict us and how to better treat our diseases of civilization.)
Understanding Human Evolution: Common Misconceptions About The Scientific Theory (Don’t make these misconceptions about evolution. Always keep up to date on the newest findings.)
Restore Western Civilization ( Enough said. As usual, gold from Brett Stevens. Amerika.org should be one of the first sites you check every day.)
I guess this was my first linkfest (ala hbd chick). I will post one a week.