1200 words
FOXP2 is a so-called “gene for” language. The gene is a transcription factor—meaning that it controls the activity of other genes. Thus, changes to FOXP2 will have changes to other genes as well. Thus, the evolution of language in humans was thought to have hinged on mutations on the FOXP2 gene. Humans that have a single-point mutation in FOXP2 “have impaired speech and grammer, but not impaired language comprehension” (Mason, et al, 2018: 403). This gene is found in numerous mammals (e.g., chimpanzees, gorillas, orangutans, rhesus macaques, and mice) but none of those mammals speak. This gene, then, is expressed in the areas of the brain that affects motor functioning, which includes the coordination needed to create words.
Mice and humans at the FOXP2 gene only differ by 3 amino acids. Only one amino acid difference exists between gorillas, chimps, mice, and macaques, who all have identical amino acid sequences on FOXP2. Furthermore, two more amino acid sequences differ between humans and the sequences which is shared by chimpanzees, gorillas, and macaques. Thus, the difference of two amino acids between humans and other primates appears to have made it possible for language to evolve. Evidence exists for strong selective pressures for the two FOXP2 mutations which allow the brain, larynx, and mouth to coordinate to produce speech. These two altered amino acids may change the ability of FOXP2 transcription factor to be phosphorylated—proteins are either activated by phosphorylation or deactivated by dephosphorylation, or the reverse.
Mason et al (2018: 403) write:
Comparative genomics efforts are now extending beyond primates. A role for FOXP2 in songbird singing and vocal learning has been proposed. Mice communicate via squeaks, with lost young mice emitting high-pitched squeaks, FOXP2 mutations leave mice squeakless. For mice and songbirds, it is a stretch to claim that FOXP2 is a language gene—but it is likely needed in the neuromuscular pathway to make sounds.
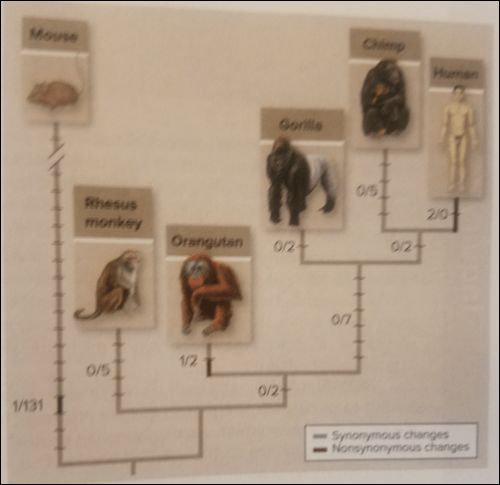
Above is Figure 18.17 from Mason et al (2018: 403). They write:
Comparisons of synonymous and nonsynonymous changes in mouse and primate FOXP2 genes indicate that changing two amino acids in the gene corresponds to the emergence of human language. Black bars represent synonymous changes; gray bars represent nonsynymous changes.
But is that the whole story? Is FOXP2 really a “gene for” language? New results call this hypothesis into question.
In their paper No Evidence for Recent Selection at FOXP2 among Diverse Human Populations, Atkinson et al (2018) did not find evidence for recent positive or balancing selection. Atksinson et al (2018) conclude that they:
do not find evidence that the FOXP2 locus or any previously implicated site within FOXP2 is associated with recent positive selection in humans. Specifically, we demonstrate that there is no evidence that the original two amino-acid substitutions were targeted by a recent sweep limited to modern humans <200 kya as suggested by Enard et al. (2002) … Any modified function of the ROI does not appear to be related to language, however, as modern southern African populations tolerate high minor allele frequencies with no apparent consequences to language faculty. We do not dispute the extensive functional evidence supporting FOXP2’s important role in the neurological processes related to language production (Lai et al., 2001, MacDermot et al., 2005, Torres-Ruiz et al., 2016). However, we show that recent natural selection in the ancestral Homo sapiens population cannot be attributed to the FOXP2 locus and thus Homo sapiens’ development of spoken language.
So the two mutations in exon 7 of FOXP2 weren’t selected and are not responsible for human language. Most likely the accelerated rate is due to loss of function (LoF) (null allele).
The gene was originally discovered in a family that had a history of speech and language disorders (Lai et al, 2001). This “speech gene” was also found in Neanderthals in 2007 (see Krasue et al, 2007). Thus, the modifications to FOXP2 occurred before humans and Neanderthals diverged.
So Atkinson et al (2018) found that the so-called sweep on FOXP2 >200KYA was a statistical artifact which was caused by lumping Africans together Caucasians and other populations. Of course, language is complicated and no one single gene will explain the emergence of human language.
This is a just-so story—that is, an ad hoc hypothesis. Humans had X, others didn’t have X or had a different form of X; therefore X explains human language faculties.
Atkinson et al’s (2018) “results represent a substantial revision to the adaptive history of FOXP2, a gene regarded as vital to human evolution.”
High evolutionary constraint among taxa but variability within Homo sapiens is compatible with a modified functional role for this locus in humans, such as a recent loss of function.
…
Therefore, this SNP must not be necessary for language function as both alleles persist at high frequency in modern human populations. Though perhaps obvious, it is important to note that there is no evidence of differences in language ability across human populations. (Atkinson et al, 2018)
This is another just-so story (Gould and Lewontin, 1976; Lloyd, 1999; Richardson, 2007; Nielsen, 2009) that seems to have bitten the dust. Of course, the functionality of FOXP2 and its role in the neurologic processes related to language; what is disputed (and refuted) is the selectionist just-so story. Selectionist explanations are necessarily ad-hoc. Thus, recent natural selection in our species cannot be attributed to FOXP2, and along with it, our language capabilities.
There is a similar objection, not for FOXP2 and selectionist hypotheses, but for the Lactase gene. Nielsen (2009) puts it succinctly:
The difference in lactose intolerance among human geographic groups, is caused by a difference in allele frequencies in and around the lactase gene (Harvey et al. 1998; Hollox et al. 2001; Enattah et al. 2002; Poulter et al. 2003). … This argument is not erected to dispute the adaptive story regarding the lactase gene, the total evidence in favor of adaptation and selection related to lactose tolerance is overwhelming in this case, but rather to argue that the combination of a functional effect and selection does not demonstrate that selection acted on the specific trait in question. … Although the presence of selection acting on genes underlying a phenotypic trait of interest does help support adaptive stories, it does not establish that selection acted directly on the specific trait of interest.
Even if there were evidence of positive selection of FOXP2 in humans, we cannot logically state that selection acted on the FOXP2 locus; functional effects and selection do not demonstrate that “selection” acted on that trait. Just-so stories (ad hoc hypotheses) “sound good”, but that’s only because they are necessarily true—one can have all the data they want, then they can think up any adaptive story to explain the data and the story will be necessarily true. Therefore, selectionist hypotheses are inherently ad hoc.
In conclusion, another selectionist hypothesis bites the dust. Nevermind the fact that, if FOXP2 were supposedly “selected-for”, there would still be the problem of free-riders (Fodor and Piattelli-Palmarini, 2010). That is, “selection” cannot “select-for” fitness-enhancing traits if/when they are coextensive with other traits—there is no way for selection to distinguish between coextensive traits and thus, it does not explain trait fixation (in this case, the fixation of FOXP2). Ad-hoc hypotheses are necessarily true—that is, they explain the data they purport to explain and only the data they purport to explain. These new results show that there is no support for positive selection at the FOXP2 locus.
RaceRealist, OK if I make a comment off-topic?
Have you seen the recent story that women have now been found to have an IQ higher than men’s by about 5 points (I may have the exact number wrong)?
I thought this was an interesting story- although I haven’t investigated it in depth. IF, however, it is the actual result of some study that is NOT actually bullshit, THEN, well, wouldn’t you say that THIS SUPPORTS YOUR OWN POINT OF VIEW (and I have to say mine as well) that IQ is a phony statistic. I mean we’ve been told for so long that IQ is immutable, and we’ve been beaten over the head with the “Bell Curve” idea that IQ averages differ by race and that those averages by race (and by gender) will not change.
Well, the story about female IQ – if it isn’t completely bogus in its own way – kind of invalidates the idea that IQ is immutable and that significant IQ differences by race are meaningful in any way.
I may even be understating the situation. You tell me.
LikeLike